Physics and Chemistry from Parsimonious Representations: Image Analysis via Invariant Variational Autoencoders
Paper and Code
Mar 30, 2023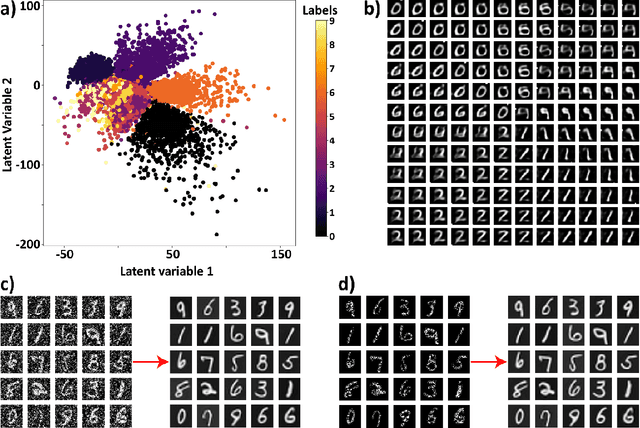
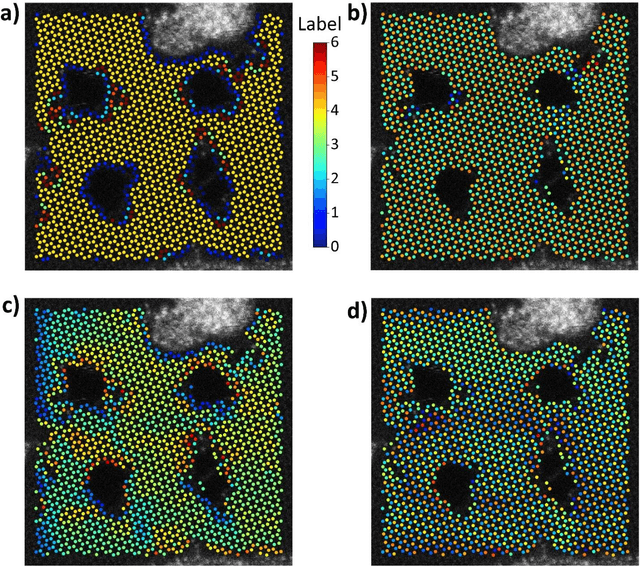
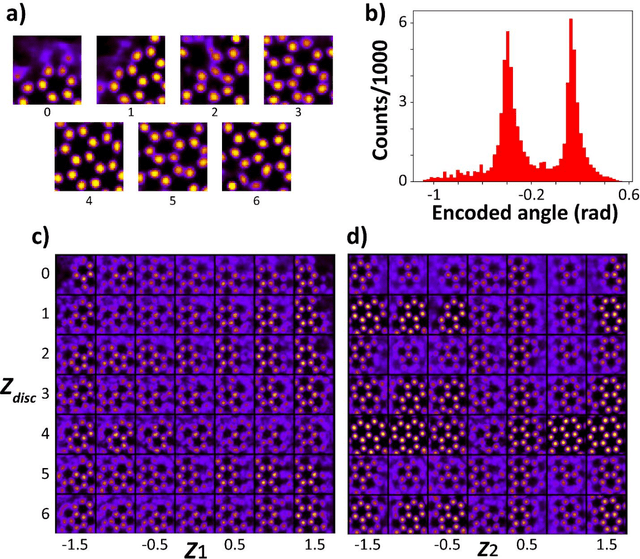

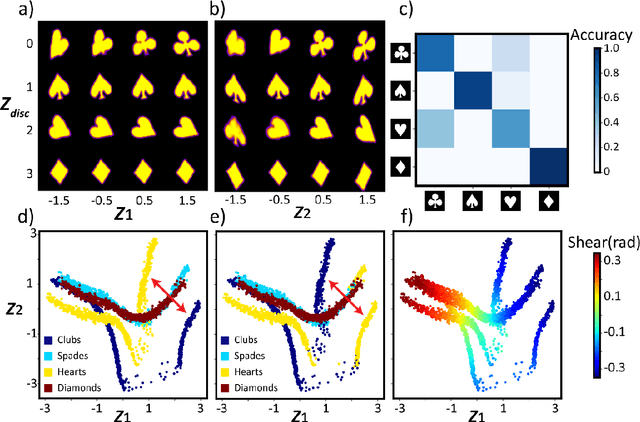
Electron, optical, and scanning probe microscopy methods are generating ever increasing volume of image data containing information on atomic and mesoscale structures and functionalities. This necessitates the development of the machine learning methods for discovery of physical and chemical phenomena from the data, such as manifestations of symmetry breaking in electron and scanning tunneling microscopy images, variability of the nanoparticles. Variational autoencoders (VAEs) are emerging as a powerful paradigm for the unsupervised data analysis, allowing to disentangle the factors of variability and discover optimal parsimonious representation. Here, we summarize recent developments in VAEs, covering the basic principles and intuition behind the VAEs. The invariant VAEs are introduced as an approach to accommodate scale and translation invariances present in imaging data and separate known factors of variations from the ones to be discovered. We further describe the opportunities enabled by the control over VAE architecture, including conditional, semi-supervised, and joint VAEs. Several case studies of VAE applications for toy models and experimental data sets in Scanning Transmission Electron Microscopy are discussed, emphasizing the deep connection between VAE and basic physical principles. All the codes used here are available at https://github.com/saimani5/VAE-tutorials and this article can be used as an application guide when applying these to own data sets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge