Patch-Based Encoder-Decoder Architecture for Automatic Transmitted Light to Fluorescence Imaging Transition: Contribution to the LightMyCells Challenge
Paper and Code
Jun 03, 2024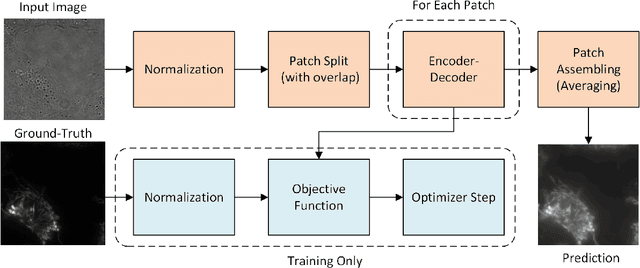
Automatic prediction of fluorescently labeled organelles from label-free transmitted light input images is an important, yet difficult task. The traditional way to obtain fluorescence images is related to performing biochemical labeling which is time-consuming and costly. Therefore, an automatic algorithm to perform the task based on the label-free transmitted light microscopy could be strongly beneficial. The importance of the task motivated researchers from the France-BioImaging to organize the LightMyCells challenge where the goal is to propose an algorithm that automatically predicts the fluorescently labeled nucleus, mitochondria, tubulin, and actin, based on the input consisting of bright field, phase contrast, or differential interference contrast microscopic images. In this work, we present the contribution of the AGHSSO team based on a carefully prepared and trained encoder-decoder deep neural network that achieves a considerable score in the challenge, being placed among the best-performing teams.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge