Multi-Objective Evolutionary approach for the Performance Improvement of Learners using Ensembling Feature selection and Discretization Technique on Medical data
Paper and Code
Apr 16, 2020
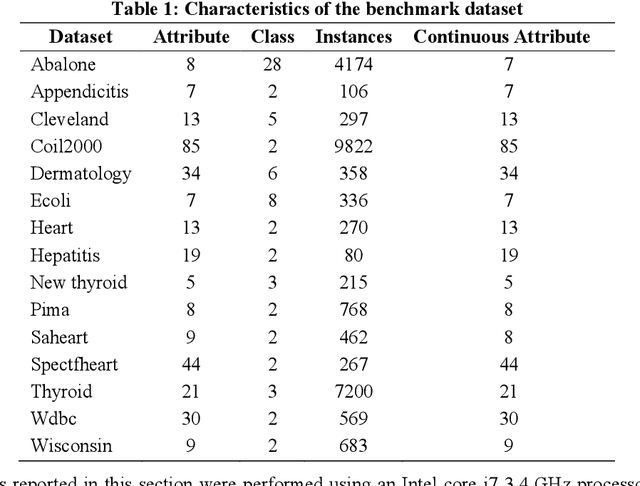
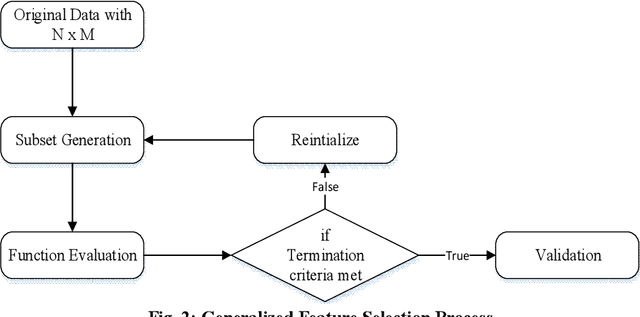
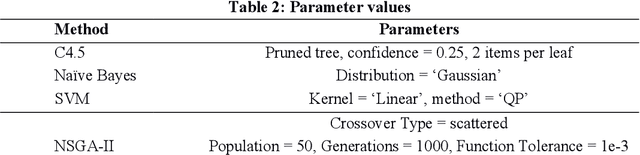
Biomedical data is filled with continuous real values; these values in the feature set tend to create problems like underfitting, the curse of dimensionality and increase in misclassification rate because of higher variance. In response, pre-processing techniques on dataset minimizes the side effects and have shown success in maintaining the adequate accuracy. Feature selection and discretization are the two necessary preprocessing steps that were effectively employed to handle the data redundancies in the biomedical data. However, in the previous works, the absence of unified effort by integrating feature selection and discretization together in solving the data redundancy problem leads to the disjoint and fragmented field. This paper proposes a novel multi-objective based dimensionality reduction framework, which incorporates both discretization and feature reduction as an ensemble model for performing feature selection and discretization. Selection of optimal features and the categorization of discretized and non-discretized features from the feature subset is governed by the multi-objective genetic algorithm (NSGA-II). The two objective, minimizing the error rate during the feature selection and maximizing the information gain while discretization is considered as fitness criteria.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge