Marginalization in Bayesian Networks: Integrating Exact and Approximate Inference
Paper and Code
Dec 16, 2021
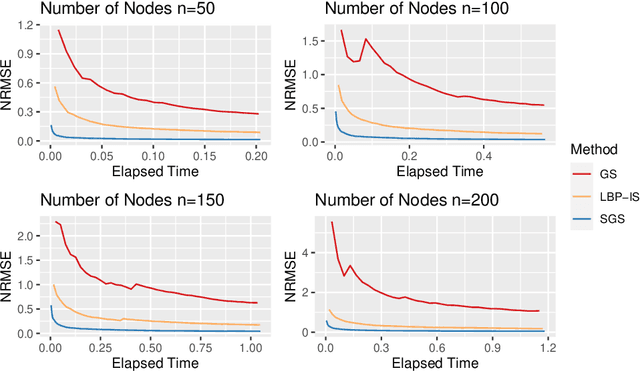
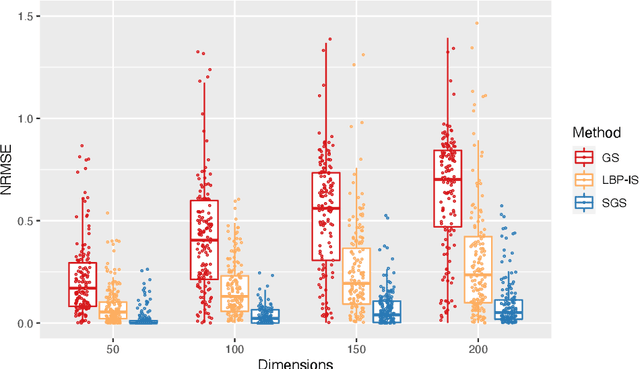
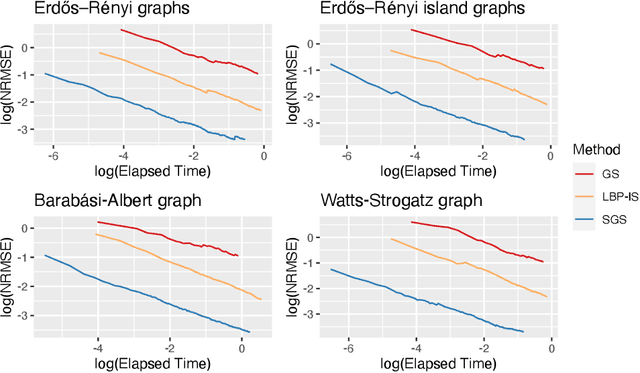
Bayesian Networks are probabilistic graphical models that can compactly represent dependencies among random variables. Missing data and hidden variables require calculating the marginal probability distribution of a subset of the variables. While knowledge of the marginal probability distribution is crucial for various problems in statistics and machine learning, its exact computation is generally not feasible for categorical variables due to the NP-hardness of this task. We develop a divide-and-conquer approach using the graphical properties of Bayesian networks to split the computation of the marginal probability distribution into sub-calculations of lower dimensionality, reducing the overall computational complexity. Exploiting this property, we present an efficient and scalable algorithm for estimating the marginal probability distribution for categorical variables. The novel method is compared against state-of-the-art approximate inference methods in a benchmarking study, where it displays superior performance. As an immediate application, we demonstrate how the marginal probability distribution can be used to classify incomplete data against Bayesian networks and use this approach for identifying the cancer subtype of kidney cancer patient samples.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge