Low dosage 3D volume fluorescence microscopy imaging using compressive sensing
Paper and Code
Jan 03, 2022
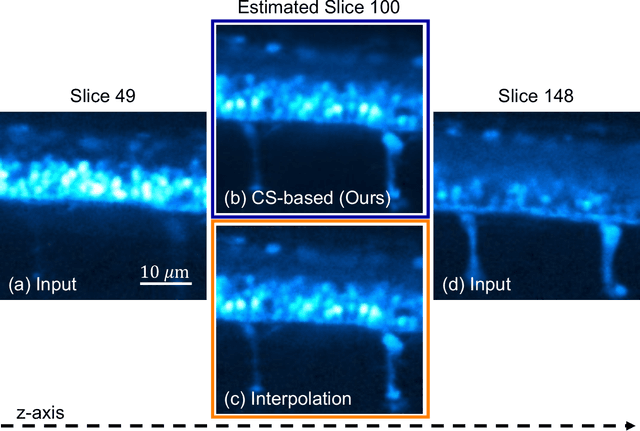
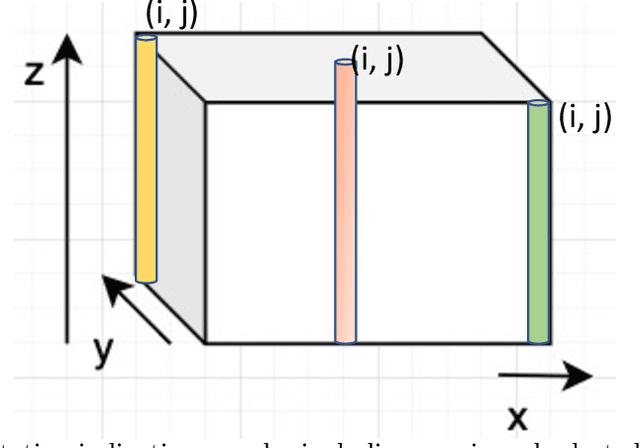
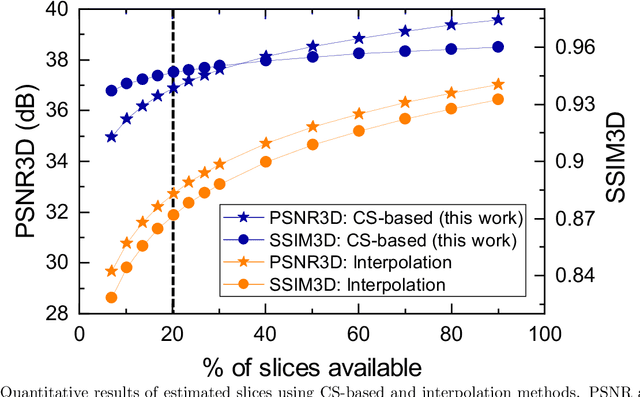

Fluorescence microscopy has been a significant tool to observe long-term imaging of embryos (in vivo) growth over time. However, cumulative exposure is phototoxic to such sensitive live samples. While techniques like light-sheet fluorescence microscopy (LSFM) allow for reduced exposure, it is not well suited for deep imaging models. Other computational techniques are computationally expensive and often lack restoration quality. To address this challenge, one can use various low-dosage imaging techniques that are developed to achieve the 3D volume reconstruction using a few slices in the axial direction (z-axis); however, they often lack restoration quality. Also, acquiring dense images (with small steps) in the axial direction is computationally expensive. To address this challenge, we present a compressive sensing (CS) based approach to fully reconstruct 3D volumes with the same signal-to-noise ratio (SNR) with less than half of the excitation dosage. We present the theory and experimentally validate the approach. To demonstrate our technique, we capture a 3D volume of the RFP labeled neurons in the zebrafish embryo spinal cord (30um thickness) with the axial sampling of 0.1um using a confocal microscope. From the results, we observe the CS-based approach achieves accurate 3D volume reconstruction from less than 20% of the entire stack optical sections. The developed CS-based methodology in this work can be easily applied to other deep imaging modalities such as two-photon and light-sheet microscopy, where reducing sample photo-toxicity is a critical challenge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge