Learning to Model the Relationship Between Brain Structural and Functional Connectomes
Paper and Code
Dec 18, 2021
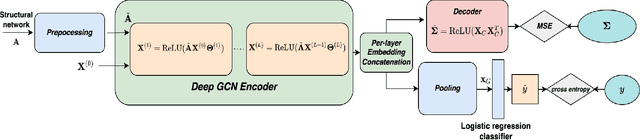


Recent advances in neuroimaging along with algorithmic innovations in statistical learning from network data offer a unique pathway to integrate brain structure and function, and thus facilitate revealing some of the brain's organizing principles at the system level. In this direction, we develop a supervised graph representation learning framework to model the relationship between brain structural connectivity (SC) and functional connectivity (FC) via a graph encoder-decoder system, where the SC is used as input to predict empirical FC. A trainable graph convolutional encoder captures direct and indirect interactions between brain regions-of-interest that mimic actual neural communications, as well as to integrate information from both the structural network topology and nodal (i.e., region-specific) attributes. The encoder learns node-level SC embeddings which are combined to generate (whole brain) graph-level representations for reconstructing empirical FC networks. The proposed end-to-end model utilizes a multi-objective loss function to jointly reconstruct FC networks and learn discriminative graph representations of the SC-to-FC mapping for downstream subject (i.e., graph-level) classification. Comprehensive experiments demonstrate that the learnt representations of said relationship capture valuable information from the intrinsic properties of the subject's brain networks and lead to improved accuracy in classifying a large population of heavy drinkers and non-drinkers from the Human Connectome Project. Our work offers new insights on the relationship between brain networks that support the promising prospect of using graph representation learning to discover more about human brain activity and function.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge