How Transferable Are Self-supervised Features in Medical Image Classification Tasks?
Paper and Code
Aug 23, 2021


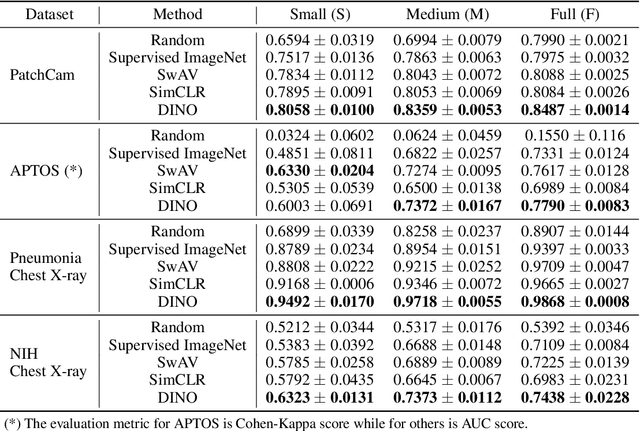
Transfer learning has become a standard practice to mitigate the lack of labeled data in medical classification tasks. Whereas finetuning a downstream task using supervised ImageNet pretrained features is straightforward and extensively investigated in many works, there is little study on the usefulness of self-supervised pretraining. In this paper, we assess the transferability of ImageNet self-supervisedpretraining by evaluating the performance of models initialized with pretrained features from three self-supervised techniques (SimCLR, SwAV, and DINO) on selected medical classification tasks. The chosen tasks cover tumor detection in sentinel axillary lymph node images, diabetic retinopathy classification in fundus images, and multiple pathological condition classification in chest X-ray images. We demonstrate that self-supervised pretrained models yield richer embeddings than their supervised counterpart, which benefits downstream tasks in view of both linear evaluation and finetuning. For example, in view of linear evaluation at acritically small subset of the data, we see an improvement up to 14.79% in Kappa score in the diabetic retinopathy classification task, 5.4% in AUC in the tumor classification task, 7.03% AUC in the pneumonia detection, and 9.4% in AUC in the detection of pathological conditions in chest X-ray. In addition, we introduce Dynamic Visual Meta-Embedding (DVME) as an end-to-end transfer learning approach that fuses pretrained embeddings from multiple models. We show that the collective representation obtained by DVME leads to a significant improvement in the performance of selected tasks compared to using a single pretrained model approach and can be generalized to any combination of pretrained models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge