Graph Based Link Prediction between Human Phenotypes and Genes
Paper and Code
Jun 01, 2021
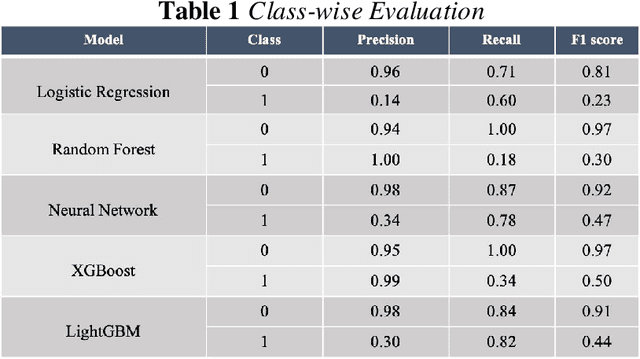
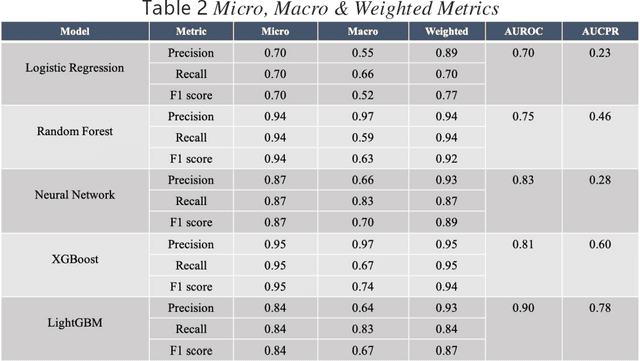
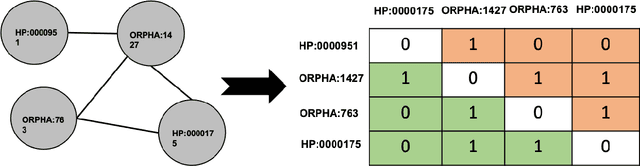
Background: The learning of genotype-phenotype associations and history of human disease by doing detailed and precise analysis of phenotypic abnormalities can be defined as deep phenotyping. To understand and detect this interaction between phenotype and genotype is a fundamental step when translating precision medicine to clinical practice. The recent advances in the field of machine learning is efficient to predict these interactions between abnormal human phenotypes and genes. Methods: In this study, we developed a framework to predict links between human phenotype ontology (HPO) and genes. The annotation data from the heterogeneous knowledge resources i.e., orphanet, is used to parse human phenotype-gene associations. To generate the embeddings for the nodes (HPO & genes), an algorithm called node2vec was used. It performs node sampling on this graph based on random walks, then learns features over these sampled nodes to generate embeddings. These embeddings were used to perform the downstream task to predict the presence of the link between these nodes using 5 different supervised machine learning algorithms. Results: The downstream link prediction task shows that the Gradient Boosting Decision Tree based model (LightGBM) achieved an optimal AUROC 0.904 and AUCPR 0.784. In addition, LightGBM achieved an optimal weighted F1 score of 0.87. Compared to the other 4 methods LightGBM is able to find more accurate interaction/link between human phenotype & gene pairs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge