Flexible co-data learning for high-dimensional prediction
Paper and Code
May 08, 2020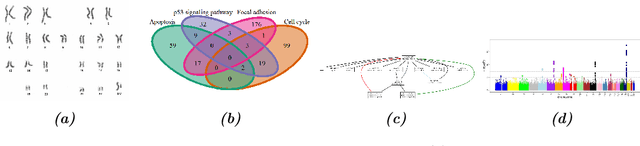

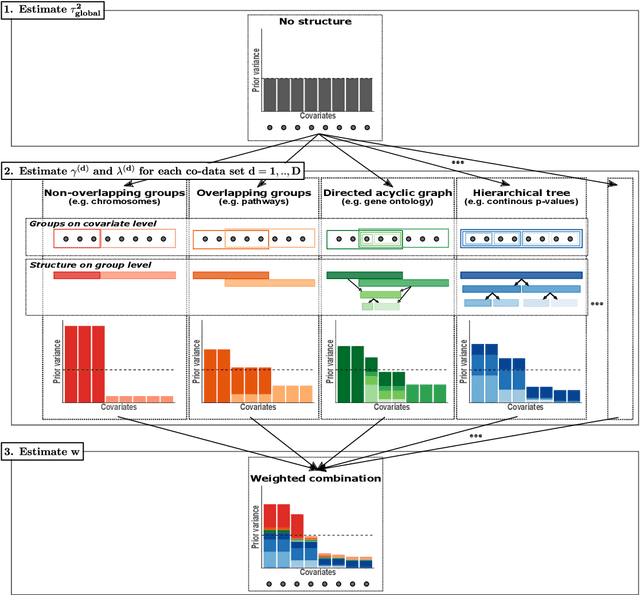
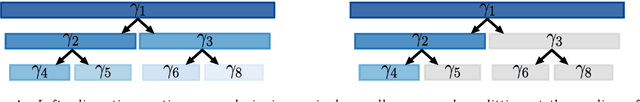
Clinical research often focuses on complex traits in which many variables play a role in mechanisms driving, or curing, diseases. Clinical prediction is hard when data is high-dimensional, but additional information, like domain knowledge and previously published studies, may be helpful to improve predictions. Such complementary data, or co-data, provide information on the covariates, such as genomic location or p-values from external studies. Our method enables exploiting multiple and various co-data sources to improve predictions. We use discrete or continuous co-data to define possibly overlapping or hierarchically structured groups of covariates. These are then used to estimate adaptive multi-group ridge penalties for generalised linear and Cox models. We combine empirical Bayes estimation of group penalty hyperparameters with an extra level of shrinkage. This renders a uniquely flexible framework as any type of shrinkage can be used on the group level. The hyperparameter shrinkage learns how relevant a specific co-data source is, counters overfitting of hyperparameters for many groups, and accounts for structured co-data. We describe various types of co-data and propose suitable forms of hypershrinkage. The method is very versatile, as it allows for integration and weighting of multiple co-data sets, inclusion of unpenalised covariates and posterior variable selection. We demonstrate it on two cancer genomics applications and show that it may improve the performance of other dense and parsimonious prognostic models substantially, and stabilises variable selection.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge