Evaluating U-net Brain Extraction for Multi-site and Longitudinal Preclinical Stroke Imaging
Paper and Code
Mar 11, 2022
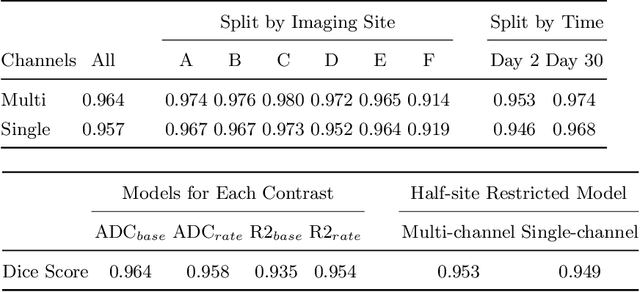
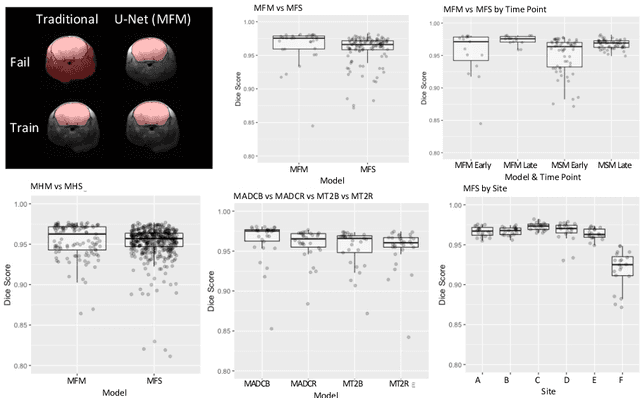
Rodent stroke models are important for evaluating treatments and understanding the pathophysiology and behavioral changes of brain ischemia, and magnetic resonance imaging (MRI) is a valuable tool for measuring outcome in preclinical studies. Brain extraction is an essential first step in most neuroimaging pipelines; however, it can be challenging in the presence of severe pathology and when dataset quality is highly variable. Convolutional neural networks (CNNs) can improve accuracy and reduce operator time, facilitating high throughput preclinical studies. As part of an ongoing preclinical stroke imaging study, we developed a deep-learning mouse brain extraction tool by using a U-net CNN. While previous studies have evaluated U-net architectures, we sought to evaluate their practical performance across data types. We ask how performance is affected with data across: six imaging centers, two time points after experimental stroke, and across four MRI contrasts. We trained, validated, and tested a typical U-net model on 240 multimodal MRI datasets including quantitative multi-echo T2 and apparent diffusivity coefficient (ADC) maps, and performed qualitative evaluation with a large preclinical stroke database (N=1,368). We describe the design and development of this system, and report our findings linking data characteristics to segmentation performance. We consistently found high accuracy and ability of the U-net architecture to generalize performance in a range of 95-97% accuracy, with only modest reductions in performance based on lower fidelity imaging hardware and brain pathology. This work can help inform the design of future preclinical rodent imaging studies and improve their scalability and reliability.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge