Disease Normalization with Graph Embeddings
Paper and Code
Oct 24, 2020
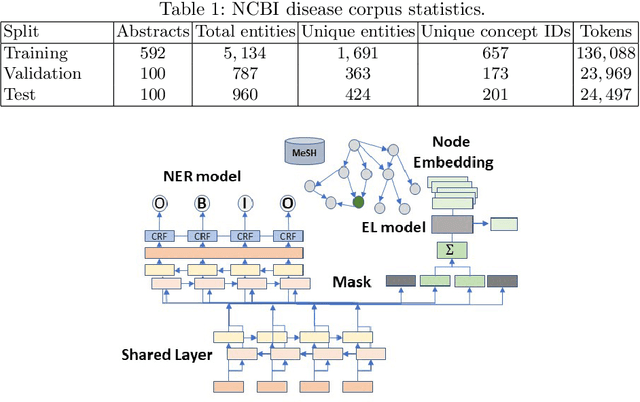
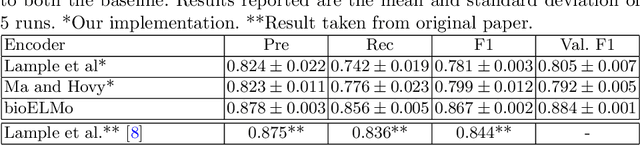
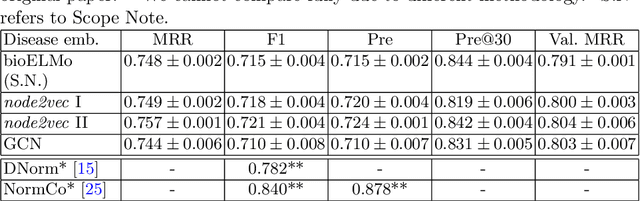
The detection and normalization of diseases in biomedical texts are key biomedical natural language processing tasks. Disease names need not only be identified, but also normalized or linked to clinical taxonomies describing diseases such as MeSH. In this paper we describe deep learning methods that tackle both tasks. We train and test our methods on the known NCBI disease benchmark corpus. We propose to represent disease names by leveraging MeSH's graphical structure together with the lexical information available in the taxonomy using graph embeddings. We also show that combining neural named entity recognition models with our graph-based entity linking methods via multitask learning leads to improved disease recognition in the NCBI corpus.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge