Discriminative protein sequence modelling with Latent Space Diffusion
Paper and Code
Mar 24, 2025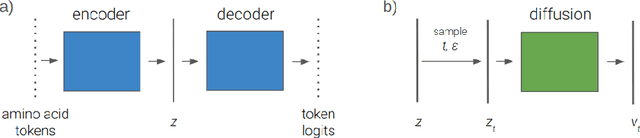



We explore a framework for protein sequence representation learning that decomposes the task between manifold learning and distributional modelling. Specifically we present a Latent Space Diffusion architecture which combines a protein sequence autoencoder with a denoising diffusion model operating on its latent space. We obtain a one-parameter family of learned representations from the diffusion model, along with the autoencoder's latent representation. We propose and evaluate two autoencoder architectures: a homogeneous model forcing amino acids of the same type to be identically distributed in the latent space, and an inhomogeneous model employing a noise-based variant of masking. As a baseline we take a latent space learned by masked language modelling, and evaluate discriminative capability on a range of protein property prediction tasks. Our finding is twofold: the diffusion models trained on both our proposed variants display higher discriminative power than the one trained on the masked language model baseline, none of the diffusion representations achieve the performance of the masked language model embeddings themselves.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge