Deep Contextual Learners for Protein Networks
Paper and Code
Jun 04, 2021
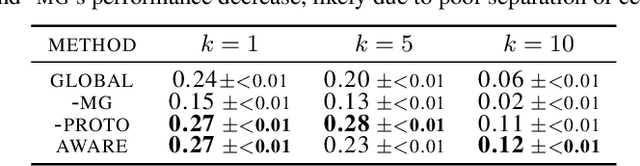

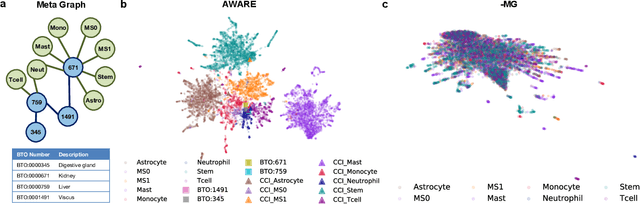
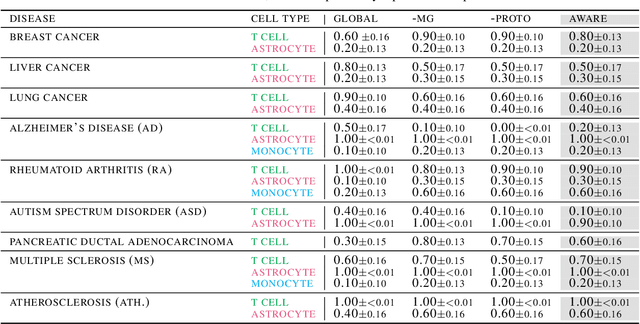
Spatial context is central to understanding health and disease. Yet reference protein interaction networks lack such contextualization, thereby limiting the study of where protein interactions likely occur in the human body. Contextualized protein interactions could better characterize genes with disease-specific interactions and elucidate diseases' manifestation in specific cell types. Here, we introduce AWARE, a graph neural message passing approach to inject cellular and tissue context into protein embeddings. AWARE optimizes for a multi-scale embedding space, whose structure reflects the topology of cell type specific networks. We construct a multi-scale network of the Human Cell Atlas and apply AWARE to learn protein, cell type, and tissue embeddings that uphold cell type and tissue hierarchies. We demonstrate AWARE on the novel task of predicting whether a gene is associated with a disease and where it most likely manifests in the human body. AWARE embeddings outperform global embeddings by at least 12.5%, highlighting the importance of contextual learners for protein networks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge