Deep Bayesian Recurrent Neural Networks for Somatic Variant Calling in Cancer
Paper and Code
Dec 06, 2019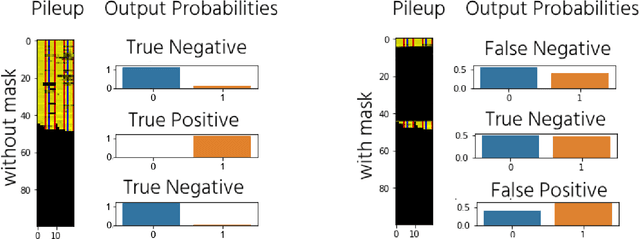
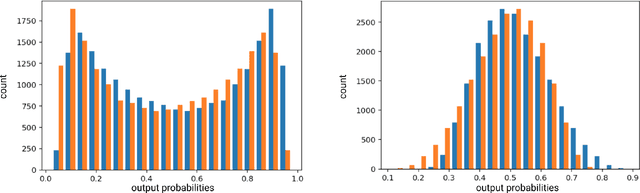
The emerging field of precision oncology relies on the accurate pinpointing of alterations in the molecular profile of a tumor to provide personalized targeted treatments. Current methodologies in the field commonly include the application of next generation sequencing technologies to a tumor sample, followed by the identification of mutations in the DNA known as somatic variants. The differentiation of these variants from sequencing error poses a classic classification problem, which has traditionally been approached with Bayesian statistics, and more recently with supervised machine learning methods such as neural networks. Although these methods provide greater accuracy, classic neural networks lack the ability to indicate the confidence of a variant call. In this paper, we explore the performance of deep Bayesian neural networks on next generation sequencing data, and their ability to give probability estimates for somatic variant calls. In addition to demonstrating similar performance in comparison to standard neural networks, we show that the resultant output probabilities make these better suited to the disparate and highly-variable sequencing data-sets these models are likely to encounter in the real world. We aim to deliver algorithms to oncologists for which model certainty better reflects accuracy, for improved clinical application. By moving away from point estimates to reliable confidence intervals, we expect the resultant clinical and treatment decisions to be more robust and more informed by the underlying reality of the tumor molecular profile.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge