Controlling the Precision-Recall Tradeoff in Differential Dependency Network Analysis
Paper and Code
Jul 09, 2013

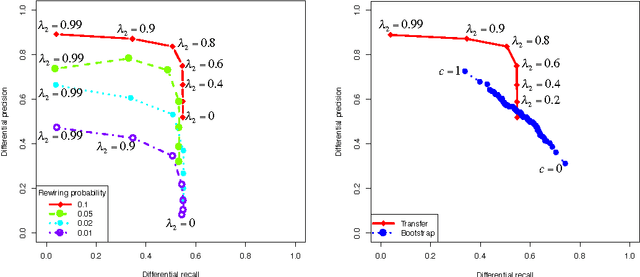
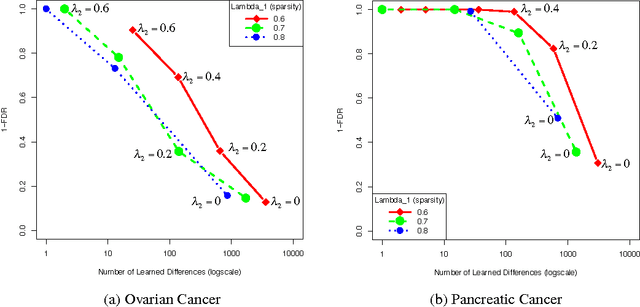
Graphical models have gained a lot of attention recently as a tool for learning and representing dependencies among variables in multivariate data. Often, domain scientists are looking specifically for differences among the dependency networks of different conditions or populations (e.g. differences between regulatory networks of different species, or differences between dependency networks of diseased versus healthy populations). The standard method for finding these differences is to learn the dependency networks for each condition independently and compare them. We show that this approach is prone to high false discovery rates (low precision) that can render the analysis useless. We then show that by imposing a bias towards learning similar dependency networks for each condition the false discovery rates can be reduced to acceptable levels, at the cost of finding a reduced number of differences. Algorithms developed in the transfer learning literature can be used to vary the strength of the imposed similarity bias and provide a natural mechanism to smoothly adjust this differential precision-recall tradeoff to cater to the requirements of the analysis conducted. We present real case studies (oncological and neurological) where domain experts use the proposed technique to extract useful differential networks that shed light on the biological processes involved in cancer and brain function.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge