Comparison of Atom Representations in Graph Neural Networks for Molecular Property Prediction
Paper and Code
Nov 23, 2020

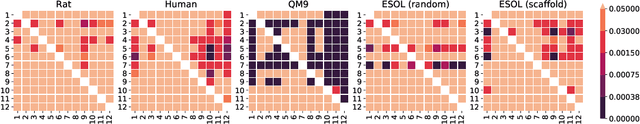

Graph neural networks have recently become a standard method for analysing chemical compounds. In the field of molecular property prediction, the emphasis is now put on designing new model architectures, and the importance of atom featurisation is oftentimes belittled. When contrasting two graph neural networks, the use of different atom features possibly leads to the incorrect attribution of the results to the network architecture. To provide a better understanding of this issue, we compare multiple atom representations for graph models and evaluate them on the prediction of free energy, solubility, and metabolic stability. To the best of our knowledge, this is the first methodological study that focuses on the relevance of atom representation to the predictive performance of graph neural networks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge