COmic: Convolutional Kernel Networks for Interpretable End-to-End Learning on (Multi-)Omics Data
Paper and Code
Dec 02, 2022
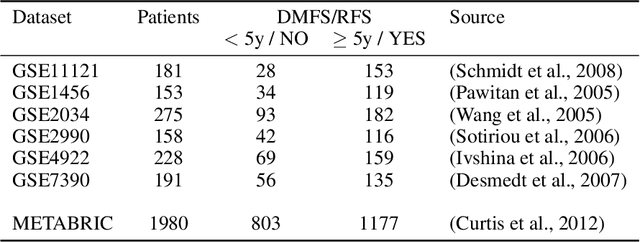
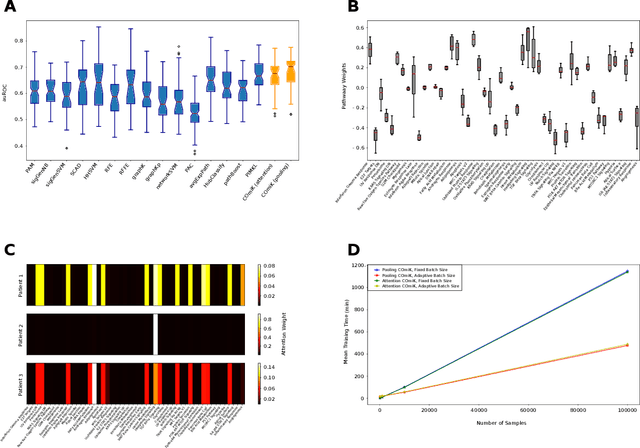
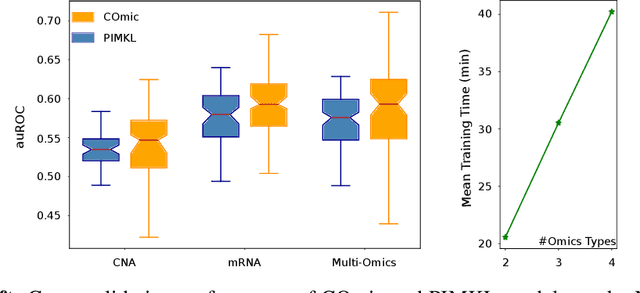
Motivation: The size of available omics datasets is steadily increasing with technological advancement in recent years. While this increase in sample size can be used to improve the performance of relevant prediction tasks in healthcare, models that are optimized for large datasets usually operate as black boxes. In high stakes scenarios, like healthcare, using a black-box model poses safety and security issues. Without an explanation about molecular factors and phenotypes that affected the prediction, healthcare providers are left with no choice but to blindly trust the models. We propose a new type of artificial neural networks, named Convolutional Omics Kernel Networks (COmic). By combining convolutional kernel networks with pathway-induced kernels, our method enables robust and interpretable end-to-end learning on omics datasets ranging in size from a few hundred to several hundreds of thousands of samples. Furthermore, COmic can be easily adapted to utilize multi-omics data. Results: We evaluate the performance capabilities of COmic on six different breast cancer cohorts. Additionally, we train COmic models on multi-omics data using the METABRIC cohort. Our models perform either better or similar to competitors on both tasks. We show how the use of pathway-induced Laplacian kernels opens the black-box nature of neural networks and results in intrinsically interpretable models that eliminate the need for \textit{post-hoc} explanation models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge