BELHD: Improving Biomedical Entity Linking with Homonoym Disambiguation
Paper and Code
Jan 10, 2024
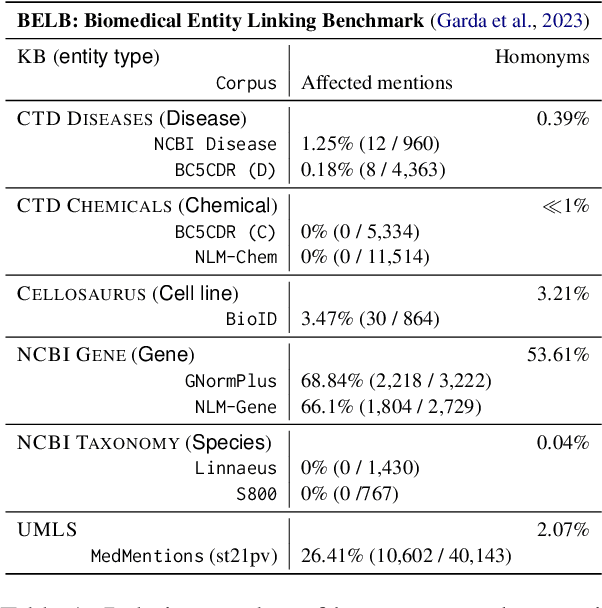
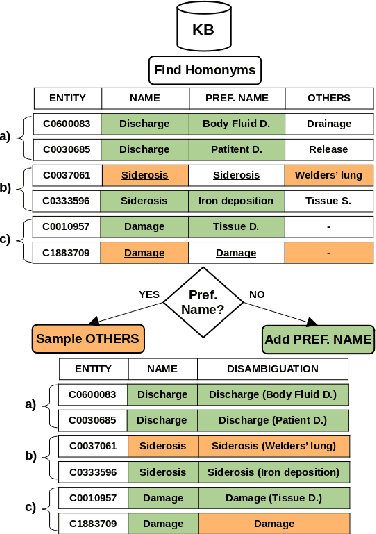
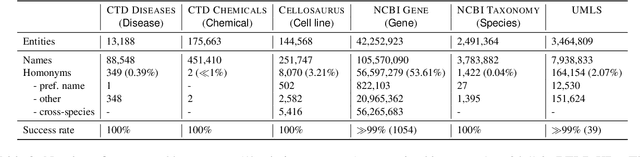
Biomedical entity linking (BEL) is the task of grounding entity mentions to a knowledge base (KB). A popular approach to the task are name-based methods, i.e. those identifying the most appropriate name in the KB for a given mention, either via dense retrieval or autoregressive modeling. However, as these methods directly return KB names, they cannot cope with homonyms, i.e. different KB entities sharing the exact same name. This significantly affects their performance, especially for KBs where homonyms account for a large amount of entity mentions (e.g. UMLS and NCBI Gene). We therefore present BELHD (Biomedical Entity Linking with Homonym Disambiguation), a new name-based method that copes with this challenge. Specifically, BELHD builds upon the BioSyn (Sung et al.,2020) model introducing two crucial extensions. First, it performs a preprocessing of the KB in which it expands homonyms with an automatically chosen disambiguating string, thus enforcing unique linking decisions. Second, we introduce candidate sharing, a novel strategy to select candidates for contrastive learning that enhances the overall training signal. Experiments with 10 corpora and five entity types show that BELHD improves upon state-of-the-art approaches, achieving the best results in 6 out 10 corpora with an average improvement of 4.55pp recall@1. Furthermore, the KB preprocessing is orthogonal to the core prediction model and thus can also improve other methods, which we exemplify for GenBioEL (Yuan et al, 2022), a generative name-based BEL approach. Code is available at: link added upon publication.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge