AutoRD: An Automatic and End-to-End System for Rare Disease Knowledge Graph Construction Based on Ontologies-enhanced Large Language Models
Paper and Code
Mar 01, 2024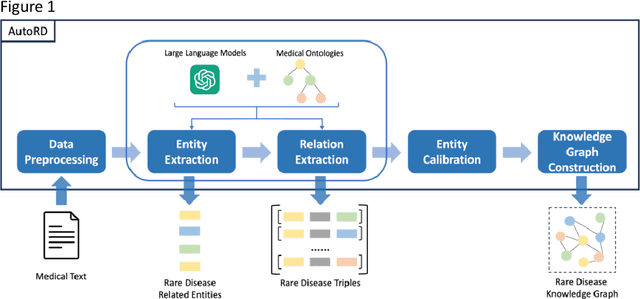


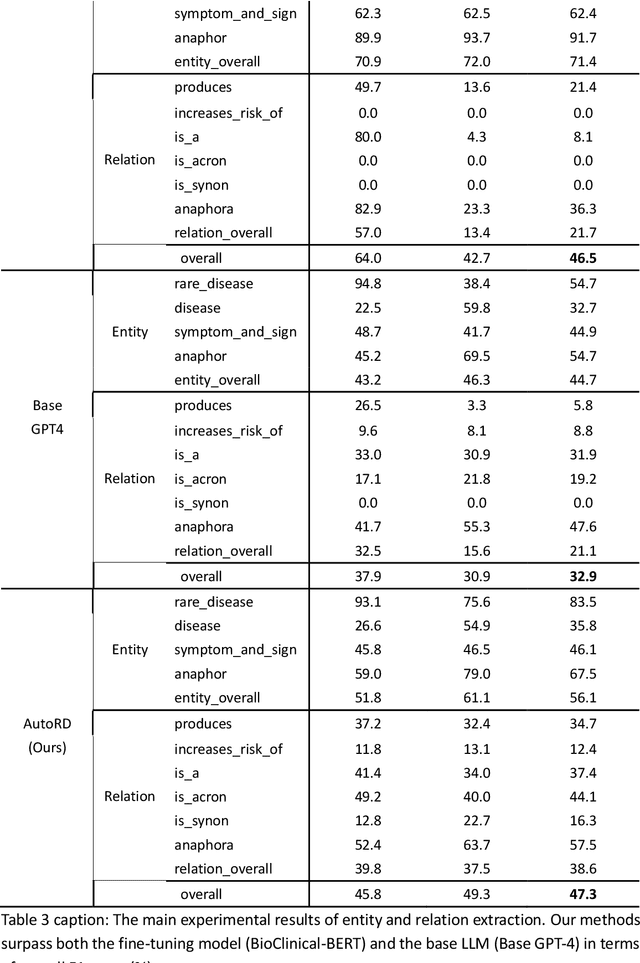
Objectives: Our objective is to create an end-to-end system called AutoRD, which automates extracting information from clinical text about rare diseases. We have conducted various tests to evaluate the performance of AutoRD and highlighted its strengths and limitations in this paper. Materials and Methods: Our system, AutoRD, is a software pipeline involving data preprocessing, entity extraction, relation extraction, entity calibration, and knowledge graph construction. We implement this using large language models and medical knowledge graphs developed from open-source medical ontologies. We quantitatively evaluate our system on entity extraction, relation extraction, and the performance of knowledge graph construction. Results: AutoRD achieves an overall F1 score of 47.3%, a 14.4% improvement compared to the base LLM. In detail, AutoRD achieves an overall entity extraction F1 score of 56.1% (rare_disease: 83.5%, disease: 35.8%, symptom_and_sign: 46.1%, anaphor: 67.5%) and an overall relation extraction F1 score of 38.6% (produces: 34.7%, increases_risk_of: 12.4%, is_a: 37.4%, is_acronym: 44.1%, is_synonym: 16.3%, anaphora: 57.5%). Our qualitative experiment also demonstrates that the performance in constructing the knowledge graph is commendable. Discussion: AutoRD demonstrates the potential of LLM applications in rare disease detection. This improvement is attributed to several design, including the integration of ontologies-enhanced LLMs. Conclusion: AutoRD is an automated end-to-end system for extracting rare disease information from text to build knowledge graphs. It uses ontologies-enhanced LLMs for a robust medical knowledge base. The superior performance of AutoRD is validated by experimental evaluations, demonstrating the potential of LLMs in healthcare.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge