Automated Detection of Acute Promyelocytic Leukemia in Blood Films and Bone Marrow Aspirates with Annotation-free Deep Learning
Paper and Code
Mar 20, 2022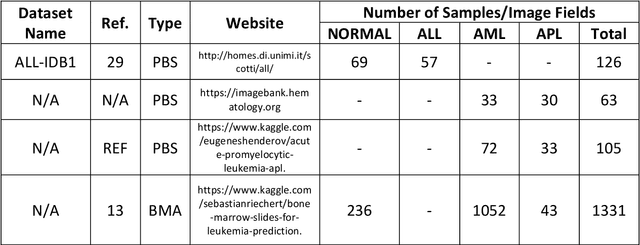
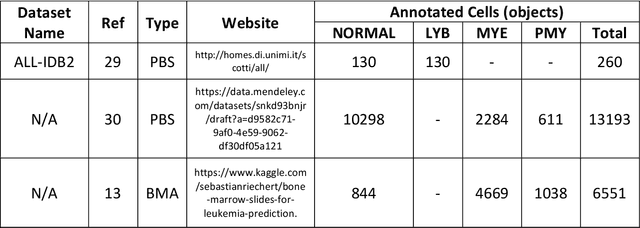
While optical microscopy inspection of blood films and bone marrow aspirates by a hematologist is a crucial step in establishing diagnosis of acute leukemia, especially in low-resource settings where other diagnostic modalities might not be available, the task remains time-consuming and prone to human inconsistencies. This has an impact especially in cases of Acute Promyelocytic Leukemia (APL) that require urgent treatment. Integration of automated computational hematopathology into clinical workflows can improve the throughput of these services and reduce cognitive human error. However, a major bottleneck in deploying such systems is a lack of sufficient cell morphological object-labels annotations to train deep learning models. We overcome this by leveraging patient diagnostic labels to train weakly-supervised models that detect different types of acute leukemia. We introduce a deep learning approach, Multiple Instance Learning for Leukocyte Identification (MILLIE), able to perform automated reliable analysis of blood films with minimal supervision. Without being trained to classify individual cells, MILLIE differentiates between acute lymphoblastic and myeloblastic leukemia in blood films. More importantly, MILLIE detects APL in blood films (AUC 0.94+/-0.04) and in bone marrow aspirates (AUC 0.99+/-0.01). MILLIE is a viable solution to augment the throughput of clinical pathways that require assessment of blood film microscopy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge