Analyzing Single Cell RNA Sequencing with Topological Nonnegative Matrix Factorization
Paper and Code
Oct 24, 2023

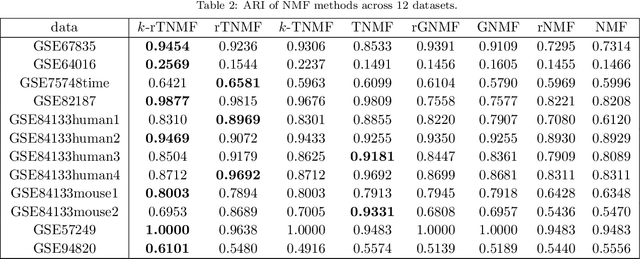

Single-cell RNA sequencing (scRNA-seq) is a relatively new technology that has stimulated enormous interest in statistics, data science, and computational biology due to the high dimensionality, complexity, and large scale associated with scRNA-seq data. Nonnegative matrix factorization (NMF) offers a unique approach due to its meta-gene interpretation of resulting low-dimensional components. However, NMF approaches suffer from the lack of multiscale analysis. This work introduces two persistent Laplacian regularized NMF methods, namely, topological NMF (TNMF) and robust topological NMF (rTNMF). By employing a total of 12 datasets, we demonstrate that the proposed TNMF and rTNMF significantly outperform all other NMF-based methods. We have also utilized TNMF and rTNMF for the visualization of popular Uniform Manifold Approximation and Projection (UMAP) and t-distributed stochastic neighbor embedding (t-SNE).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge