A novel unsupervised covid lung lesion segmentation based on the lung tissue identification
Paper and Code
Feb 24, 2022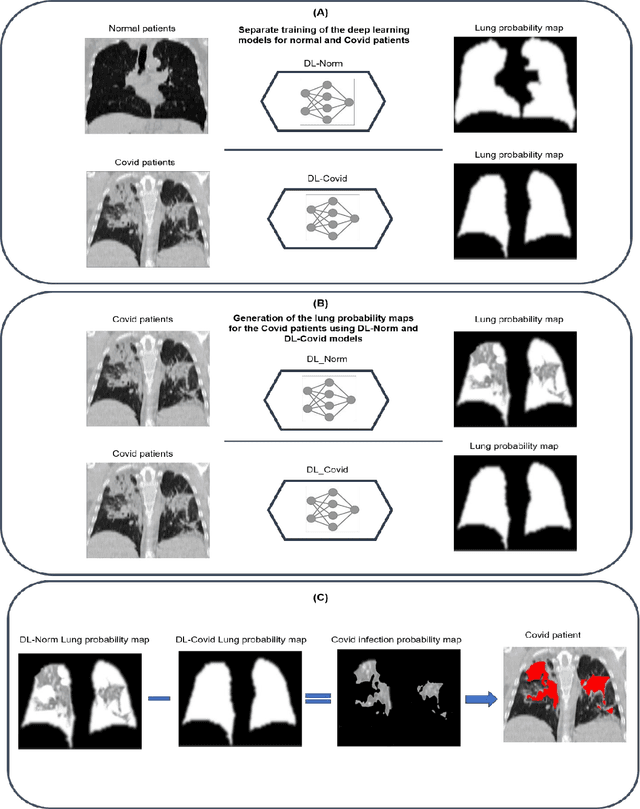
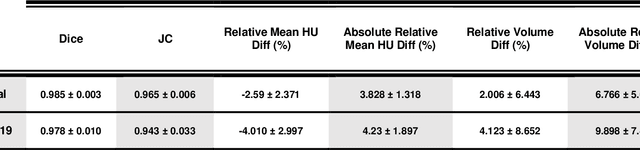
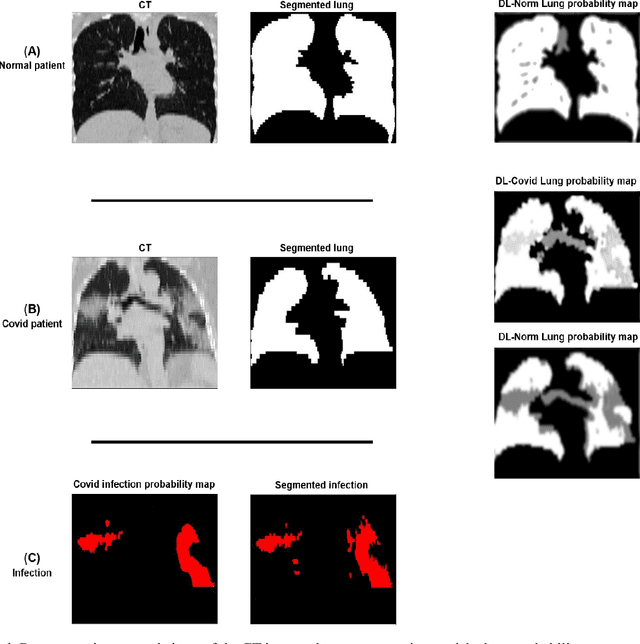

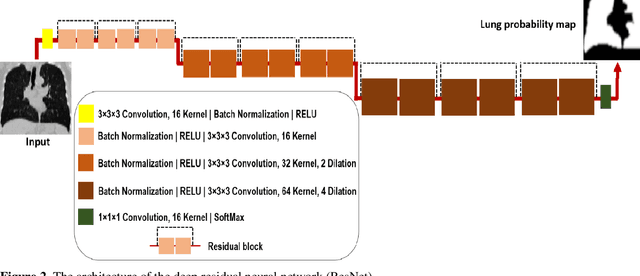
This study aimed to evaluate the performance of a novel unsupervised deep learning-based framework for automated infections lesion segmentation from CT images of Covid patients. In the first step, two residual networks were independently trained to identify the lung tissue for normal and Covid patients in a supervised manner. These two models, referred to as DL-Covid and DL-Norm for Covid-19 and normal patients, respectively, generate the voxel-wise probability maps for lung tissue identification. To detect Covid lesions, the CT image of the Covid patient is processed by the DL-Covid and DL-Norm models to obtain two lung probability maps. Since the DL-Norm model is not familiar with Covid infections within the lung, this model would assign lower probabilities to the lesions than the DL-Covid. Hence, the probability maps of the Covid infections could be generated through the subtraction of the two lung probability maps obtained from the DL-Covid and DL-Norm models. Manual lesion segmentation of 50 Covid-19 CT images was used to assess the accuracy of the unsupervised lesion segmentation approach. The Dice coefficients of 0.985 and 0.978 were achieved for the lung segmentation of normal and Covid patients in the external validation dataset, respectively. Quantitative results of infection segmentation by the proposed unsupervised method showed the Dice coefficient and Jaccard index of 0.67 and 0.60, respectively. Quantitative evaluation of the proposed unsupervised approach for Covid-19 infectious lesion segmentation showed relatively satisfactory results. Since this framework does not require any annotated dataset, it could be used to generate very large training samples for the supervised machine learning algorithms dedicated to noisy and/or weakly annotated datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge