A deep learning pipeline for cross-sectional and longitudinal multiview data integration
Paper and Code
Dec 02, 2023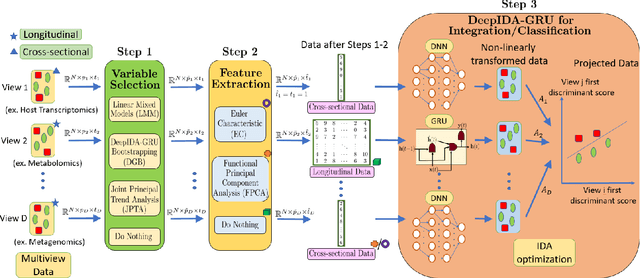



Biomedical research now commonly integrates diverse data types or views from the same individuals to better understand the pathobiology of complex diseases, but the challenge lies in meaningfully integrating these diverse views. Existing methods often require the same type of data from all views (cross-sectional data only or longitudinal data only) or do not consider any class outcome in the integration method, presenting limitations. To overcome these limitations, we have developed a pipeline that harnesses the power of statistical and deep learning methods to integrate cross-sectional and longitudinal data from multiple sources. Additionally, it identifies key variables contributing to the association between views and the separation among classes, providing deeper biological insights. This pipeline includes variable selection/ranking using linear and nonlinear methods, feature extraction using functional principal component analysis and Euler characteristics, and joint integration and classification using dense feed-forward networks and recurrent neural networks. We applied this pipeline to cross-sectional and longitudinal multi-omics data (metagenomics, transcriptomics, and metabolomics) from an inflammatory bowel disease (IBD) study and we identified microbial pathways, metabolites, and genes that discriminate by IBD status, providing information on the etiology of IBD. We conducted simulations to compare the two feature extraction methods. The proposed pipeline is available from the following GitHub repository: https://github.com/lasandrall/DeepIDA-GRU.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge