Yoga Sri Varshan V
Detection of Under-represented Samples Using Dynamic Batch Training for Brain Tumor Segmentation from MR Images
Aug 21, 2024
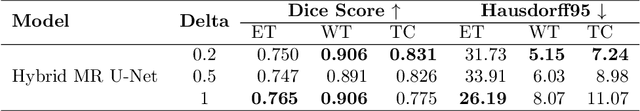


Abstract:Brain tumors in magnetic resonance imaging (MR) are difficult, time-consuming, and prone to human error. These challenges can be resolved by developing automatic brain tumor segmentation methods from MR images. Various deep-learning models based on the U-Net have been proposed for the task. These deep-learning models are trained on a dataset of tumor images and then used for segmenting the masks. Mini-batch training is a widely used method in deep learning for training. However, one of the significant challenges associated with this approach is that if the training dataset has under-represented samples or samples with complex latent representations, the model may not generalize well to these samples. The issue leads to skewed learning of the data, where the model learns to fit towards the majority representations while underestimating the under-represented samples. The proposed dynamic batch training method addresses the challenges posed by under-represented data points, data points with complex latent representation, and imbalances within the class, where some samples may be harder to learn than others. Poor performance of such samples can be identified only after the completion of the training, leading to the wastage of computational resources. Also, training easy samples after each epoch is an inefficient utilization of computation resources. To overcome these challenges, the proposed method identifies hard samples and trains such samples for more iterations compared to easier samples on the BraTS2020 dataset. Additionally, the samples trained multiple times are identified and it provides a way to identify hard samples in the BraTS2020 dataset. The comparison of the proposed training approach with U-Net and other models in the literature highlights the capabilities of the proposed training approach.
Integrating Edge Information into Ground Truth for the Segmentation of the Optic Disc and Cup from Fundus Images
Aug 09, 2024Abstract:Optic disc and cup segmentation helps in the diagnosis of glaucoma, myocardial infarction, and diabetic retinopathy. Most deep learning methods developed to perform segmentation tasks are built on top of a U-Net-based model architecture. Nevertheless, U-Net and its variants have a tendency to over-segment/ under-segment the required regions of interest. Since the most important outcome is the value of cup-to-disc ratio and not the segmented regions themselves, we are more concerned about the boundaries rather than the regions under the boundaries. This makes learning edges important as compared to learning the regions. In the proposed work, the authors aim to extract both edges of the optic disc and cup from the ground truth using a Laplacian filter. Next, edges are reconstructed to obtain an edge ground truth in addition to the optic disc-cup ground truth. Utilizing both ground truths, the authors study several U-Net and its variant architectures with and without optic disc and cup edges as target, along with the optic disc-cup ground truth for segmentation. The authors have used the REFUGE benchmark dataset and the Drishti-GS dataset to perform the study, and the results are tabulated for the dice and the Hausdorff distance metrics. In the case of the REFUGE dataset, the optic disc mean dice score has improved from 0.7425 to 0.8859 while the mean Hausdorff distance has reduced from 6.5810 to 3.0540 for the baseline U-Net model. Similarly, the optic cup mean dice score has improved from 0.6970 to 0.8639 while the mean Hausdorff distance has reduced from 5.2340 to 2.6323 for the same model. Similar improvement has been observed for the Drishti-GS dataset as well. Compared to the baseline U-Net and its variants (i.e) the Attention U-Net and the U-Net++, the models that learn integrated edges along with the optic disc and cup regions performed well in both validation and testing datasets.
Explainable AI: Comparative Analysis of Normal and Dilated ResNet Models for Fundus Disease Classification
Jul 07, 2024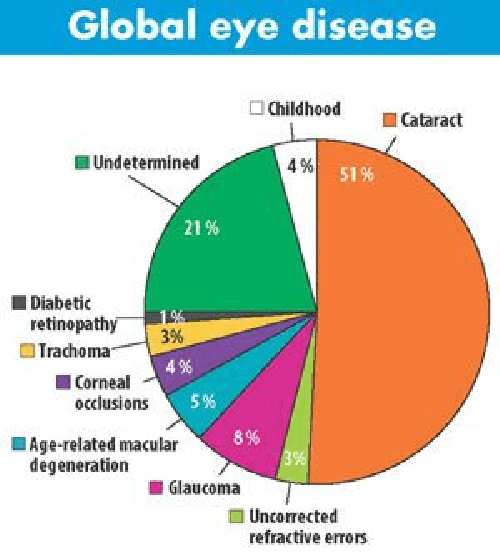



Abstract:This paper presents dilated Residual Network (ResNet) models for disease classification from retinal fundus images. Dilated convolution filters are used to replace normal convolution filters in the higher layers of the ResNet model (dilated ResNet) in order to improve the receptive field compared to the normal ResNet model for disease classification. This study introduces computer-assisted diagnostic tools that employ deep learning, enhanced with explainable AI techniques. These techniques aim to make the tool's decision-making process transparent, thereby enabling medical professionals to understand and trust the AI's diagnostic decision. They are particularly relevant in today's healthcare landscape, where there is a growing demand for transparency in AI applications to ensure their reliability and ethical use. The dilated ResNet is used as a replacement for the normal ResNet to enhance the classification accuracy of retinal eye diseases and reduce the required computing time. The dataset used in this work is the Ocular Disease Intelligent Recognition (ODIR) dataset which is a structured ophthalmic database with eight classes covering most of the common retinal eye diseases. The evaluation metrics used in this work include precision, recall, accuracy, and F1 score. In this work, a comparative study has been made between normal ResNet models and dilated ResNet models on five variants namely ResNet-18, ResNet-34, ResNet-50, ResNet-101, and ResNet-152. The dilated ResNet model shows promising results as compared to normal ResNet with an average F1 score of 0.71, 0.70, 0.69, 0.67, and 0.70 respectively for the above respective variants in ODIR multiclass disease classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge