Yair Ben Sahel
Self-STORM: Deep Unrolled Self-Supervised Learning for Super-Resolution Microscopy
Mar 25, 2024



Abstract:The use of fluorescent molecules to create long sequences of low-density, diffraction-limited images enables highly-precise molecule localization. However, this methodology requires lengthy imaging times, which limits the ability to view dynamic interactions of live cells on short time scales. Many techniques have been developed to reduce the number of frames needed for localization, from classic iterative optimization to deep neural networks. Particularly, deep algorithm unrolling utilizes both the structure of iterative sparse recovery algorithms and the performance gains of supervised deep learning. However, the robustness of this approach is highly dependant on having sufficient training data. In this paper we introduce deep unrolled self-supervised learning, which alleviates the need for such data by training a sequence-specific, model-based autoencoder that learns only from given measurements. Our proposed method exceeds the performance of its supervised counterparts, thus allowing for robust, dynamic imaging well below the diffraction limit without any labeled training samples. Furthermore, the suggested model-based autoencoder scheme can be utilized to enhance generalization in any sparse recovery framework, without the need for external training data.
Deep Unrolled Recovery in Sparse Biological Imaging
Sep 28, 2021
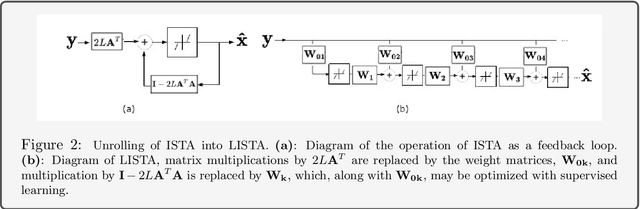

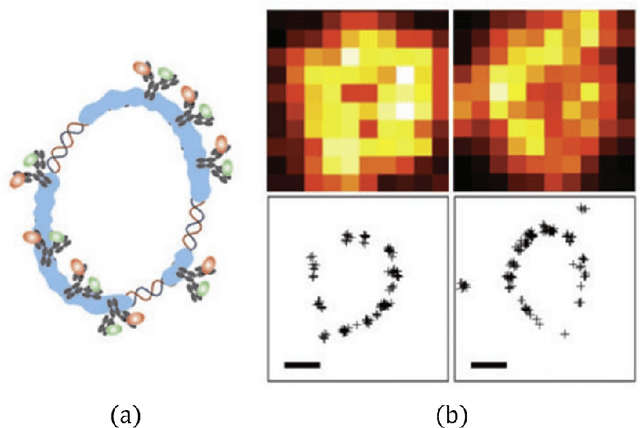

Abstract:Deep algorithm unrolling has emerged as a powerful model-based approach to develop deep architectures that combine the interpretability of iterative algorithms with the performance gains of supervised deep learning, especially in cases of sparse optimization. This framework is well-suited to applications in biological imaging, where physics-based models exist to describe the measurement process and the information to be recovered is often highly structured. Here, we review the method of deep unrolling, and show how it improves source localization in several biological imaging settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge