Vishal Srivastava
Quantifying Automation Risk in High-Automation AI Systems: A Bayesian Framework for Failure Propagation and Optimal Oversight
Feb 22, 2026Abstract:Organizations across finance, healthcare, transportation, content moderation, and critical infrastructure are rapidly deploying highly automated AI systems, yet they lack principled methods to quantify how increasing automation amplifies harm when failures occur. We propose a parsimonious Bayesian risk decomposition expressing expected loss as the product of three terms: the probability of system failure, the conditional probability that a failure propagates into harm given the automation level, and the expected severity of harm. This framework isolates a critical quantity -- the conditional probability that failures propagate into harm -- which captures execution and oversight risk rather than model accuracy alone. We develop complete theoretical foundations: formal proofs of the decomposition, a harm propagation equivalence theorem linking the harm propagation probability to observable execution controls, risk elasticity measures, efficient frontier analysis for automation policy, and optimal resource allocation principles with second-order conditions. We motivate the framework with an illustrative case study of the 2012 Knight Capital incident ($440M loss) as one instantiation of a broadly applicable failure pattern, and characterize the research design required to empirically validate the framework at scale across deployment domains. This work provides the theoretical foundations for a new class of deployment-focused risk governance tools for agentic and automated AI systems.
Fundamental Limits of Black-Box Safety Evaluation: Information-Theoretic and Computational Barriers from Latent Context Conditioning
Feb 19, 2026Abstract:Black-box safety evaluation of AI systems assumes model behavior on test distributions reliably predicts deployment performance. We formalize and challenge this assumption through latent context-conditioned policies -- models whose outputs depend on unobserved internal variables that are rare under evaluation but prevalent under deployment. We establish fundamental limits showing that no black-box evaluator can reliably estimate deployment risk for such models. (1) Passive evaluation: For evaluators sampling i.i.d. from D_eval, we prove minimax lower bounds via Le Cam's method: any estimator incurs expected absolute error >= (5/24)*delta*L approximately 0.208*delta*L, where delta is trigger probability under deployment and L is the loss gap. (2) Adaptive evaluation: Using a hash-based trigger construction and Yao's minimax principle, worst-case error remains >= delta*L/16 even for fully adaptive querying when D_dep is supported over a sufficiently large domain; detection requires Theta(1/epsilon) queries. (3) Computational separation: Under trapdoor one-way function assumptions, deployment environments possessing privileged information can activate unsafe behaviors that any polynomial-time evaluator without the trapdoor cannot distinguish. For white-box probing, estimating deployment risk to accuracy epsilon_R requires O(1/(gamma^2 * epsilon_R^2)) samples, where gamma = alpha_0 + alpha_1 - 1 measures probe quality, and we provide explicit bias correction under probe error. Our results quantify when black-box testing is statistically underdetermined and provide explicit criteria for when additional safeguards -- architectural constraints, training-time guarantees, interpretability, and deployment monitoring -- are mathematically necessary for worst-case safety assurance.
Incorporating Total Variation Regularization in the design of an intelligent Query by Humming system
Feb 09, 2023



Abstract:A Query-By-Humming (QBH) system constitutes a particular case of music information retrieval where the input is a user-hummed melody and the output is the original song which contains that melody. A typical QBH system consists of melody extraction and candidate melody retrieval. For melody extraction, accurate note transcription is the key enabling technology. However, current transcription methods are unable to definitively capture the melody and address inaccuracies in user-hummed queries. In this paper, we incorporate Total Variation Regularization (TVR) to denoise queries. This approach accounts for user error in humming without loss of meaningful data and reliably captures the underlying melody. For candidate melody retrieval, we employ a deep learning approach to time series classification using a Fully Convolutional Neural Network. The trained network classifies the incoming query as belonging to one of the target songs. For our experiments, we use Roger Jang's MIR-QBSH dataset which is the standard MIREX dataset. We demonstrate that inclusion of TVR denoised queries in the training set enhances the overall accuracy of the system to 93% which is higher than other state-of-the-art QBH systems.
Self-attention based BiLSTM-CNN classifier for the prediction of ischemic and non-ischemic cardiomyopathy
Jul 29, 2019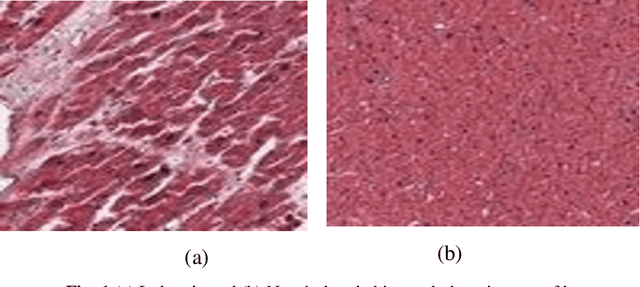
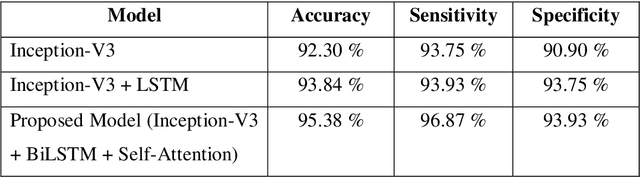
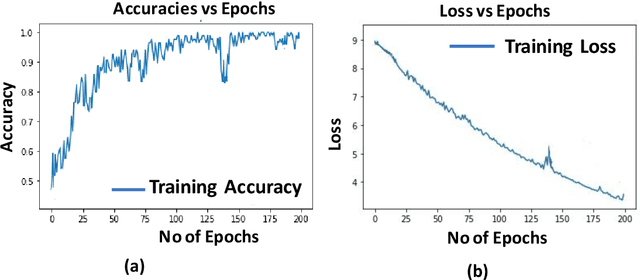
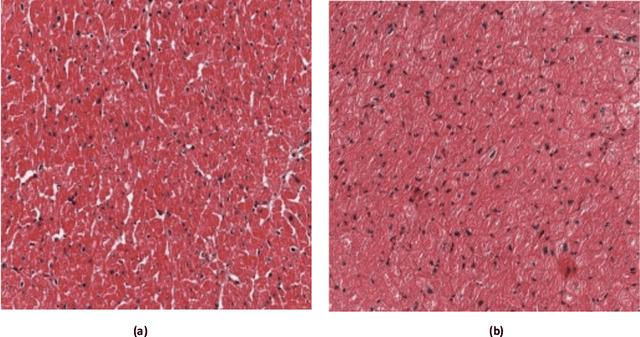
Abstract:Heart Failure is a major component of healthcare expenditure and a leading cause of mortality worldwide. Despite higher inter-rater variability, endomyocardial biopsy (EMB) is still regarded as the standard technique, used to identify the cause (e.g. ischemic or non-ischemic cardiomyopathy, coronary artery disease, myocardial infarction etc.) of unexplained heart failure. In this paper, we focus on identifying cardiomyopathy as ischemic or non-ischemic. For this, we propose and implement a new unified architecture comprising CNN (inception-V3 model) and bidirectional LSTM (BiLSTM) with self-attention mechanism to predict the ischemic or non-ischemic to classify cardiomyopathy using histopathological images. The proposed model is based on self-attention that implicitly focuses on the information outputted from the hidden layers of BiLSTM. Through our results we demonstrate that this framework carries a high learning capacity and is able to improve the classification performance.
Deep learning enabled multi-wavelength spatial coherence microscope for the classification of malaria-infected stages with limited labelled data size
Mar 14, 2019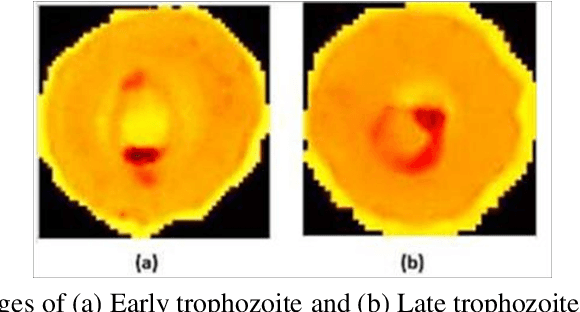
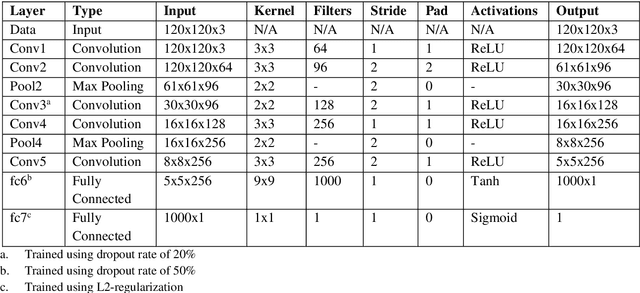
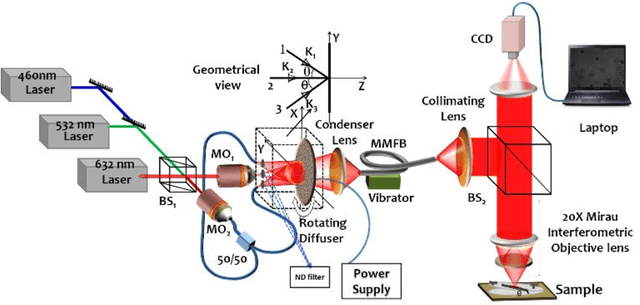
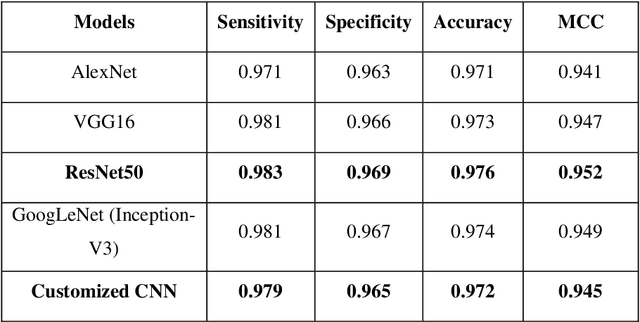
Abstract:Malaria is a life-threatening mosquito-borne blood disease, hence early detection is very crucial for health. The conventional method for the detection is a microscopic examination of Giemsa-stained blood smears, which needs a highly trained skilled technician. Automated classifications of different stages of malaria still a challenging task, especially having poor sensitivity in detecting the early trophozoite and late trophozoite or schizont stage with limited labelled datasize. The study aims to develop a fast, robust and fully automated system for the classification of different stages of malaria with limited data size by using the pre-trained convolutional neural networks (CNNs) as a classifier and multi-wavelength to increase the sample size. We also compare our customized CNN with other well-known CNNs and shows that our network have a comparable performance with less computational time. We believe that our proposed method can be applied to other limited labelled biological datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge