Sunil Mathew
Generation of patient specific cardiac chamber models using generative neural networks under a Bayesian framework for electroanatomical mapping
Nov 27, 2023Abstract:Electroanatomical mapping is a technique used in cardiology to create a detailed 3D map of the electrical activity in the heart. It is useful for diagnosis, treatment planning and real time guidance in cardiac ablation procedures to treat arrhythmias like atrial fibrillation. A probabilistic machine learning model trained on a library of CT/MRI scans of the heart can be used during electroanatomical mapping to generate a patient-specific 3D model of the chamber being mapped. The use of probabilistic machine learning models under a Bayesian framework provides a way to quantify uncertainty in results and provide a natural framework of interpretability of the model. Here we introduce a Bayesian approach to surface reconstruction of cardiac chamber models from a sparse 3D point cloud data acquired during electroanatomical mapping. We show how probabilistic graphical models trained on segmented CT/MRI data can be used to generate cardiac chamber models from few acquired locations thereby reducing procedure time and x-ray exposure. We show how they provide insight into what the neural network learns from the segmented CT/MRI images used to train the network, which provides explainability to the resulting cardiac chamber models generated by the model.
Pruning a neural network using Bayesian inference
Aug 04, 2023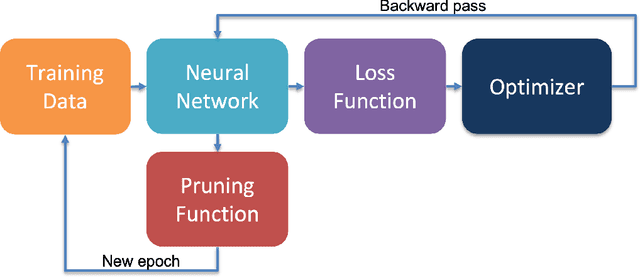



Abstract:Neural network pruning is a highly effective technique aimed at reducing the computational and memory demands of large neural networks. In this research paper, we present a novel approach to pruning neural networks utilizing Bayesian inference, which can seamlessly integrate into the training procedure. Our proposed method leverages the posterior probabilities of the neural network prior to and following pruning, enabling the calculation of Bayes factors. The calculated Bayes factors guide the iterative pruning. Through comprehensive evaluations conducted on multiple benchmarks, we demonstrate that our method achieves desired levels of sparsity while maintaining competitive accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge