Stephen M Plaza
Latent Feature Representation via Unsupervised Learning for Pattern Discovery in Massive Electron Microscopy Image Volumes
Dec 22, 2020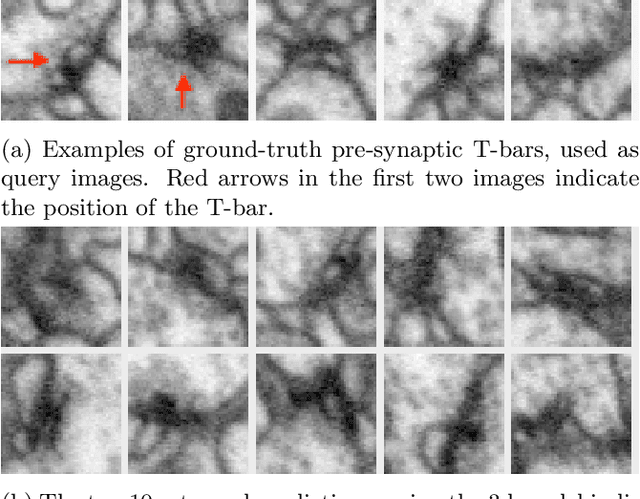
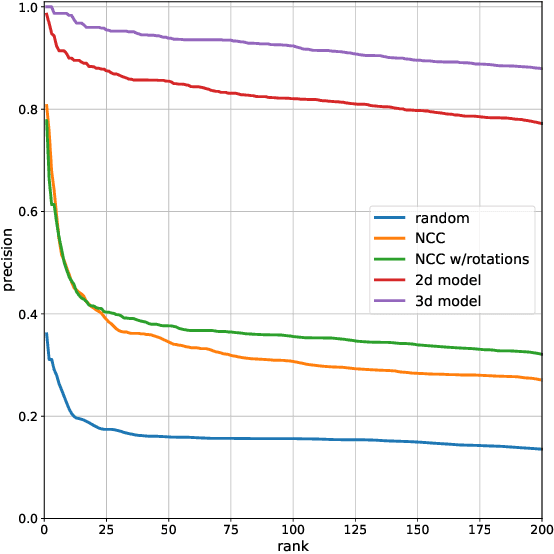

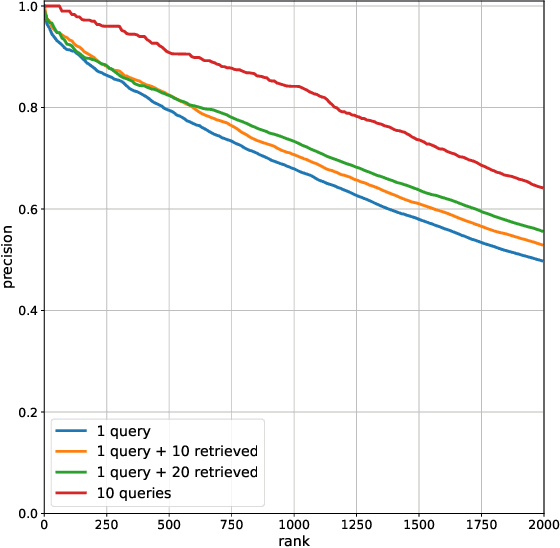
Abstract:We propose a method to facilitate exploration and analysis of new large data sets. In particular, we give an unsupervised deep learning approach to learning a latent representation that captures semantic similarity in the data set. The core idea is to use data augmentations that preserve semantic meaning to generate synthetic examples of elements whose feature representations should be close to one another. We demonstrate the utility of our method applied to nano-scale electron microscopy data, where even relatively small portions of animal brains can require terabytes of image data. Although supervised methods can be used to predict and identify known patterns of interest, the scale of the data makes it difficult to mine and analyze patterns that are not known a priori. We show the ability of our learned representation to enable query by example, so that if a scientist notices an interesting pattern in the data, they can be presented with other locations with matching patterns. We also demonstrate that clustering of data in the learned space correlates with biologically-meaningful distinctions. Finally, we introduce a visualization tool and software ecosystem to facilitate user-friendly interactive analysis and uncover interesting biological patterns. In short, our work opens possible new avenues in understanding of and discovery in large data sets, arising in domains such as EM analysis.
Automatic Neuron Type Identification by Neurite Localization in the Drosophila Medulla
Sep 05, 2014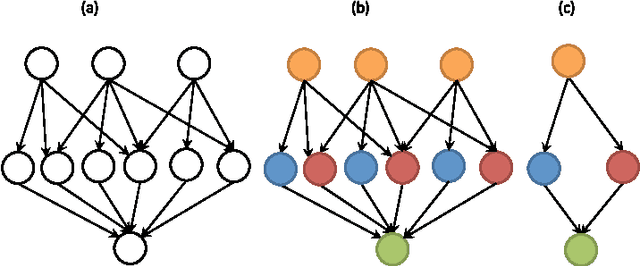
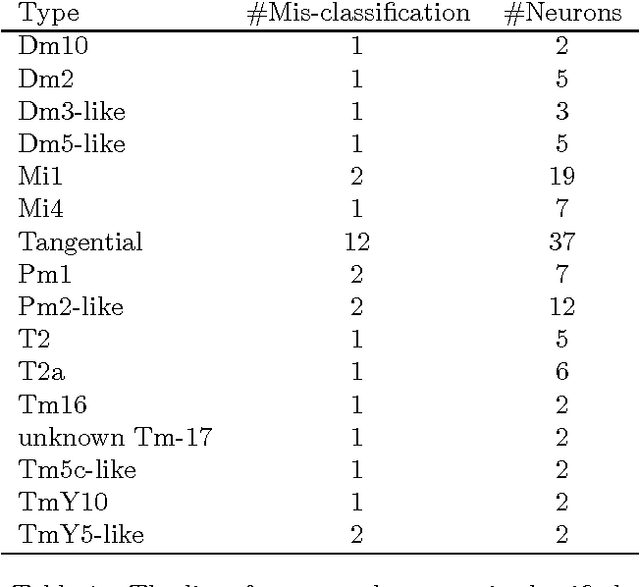
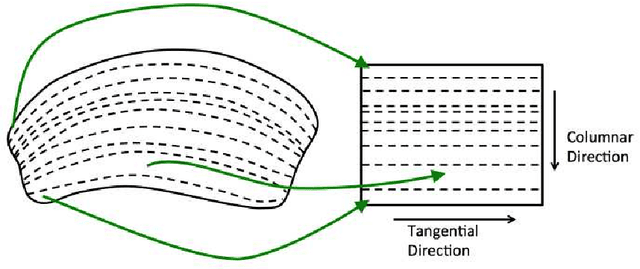
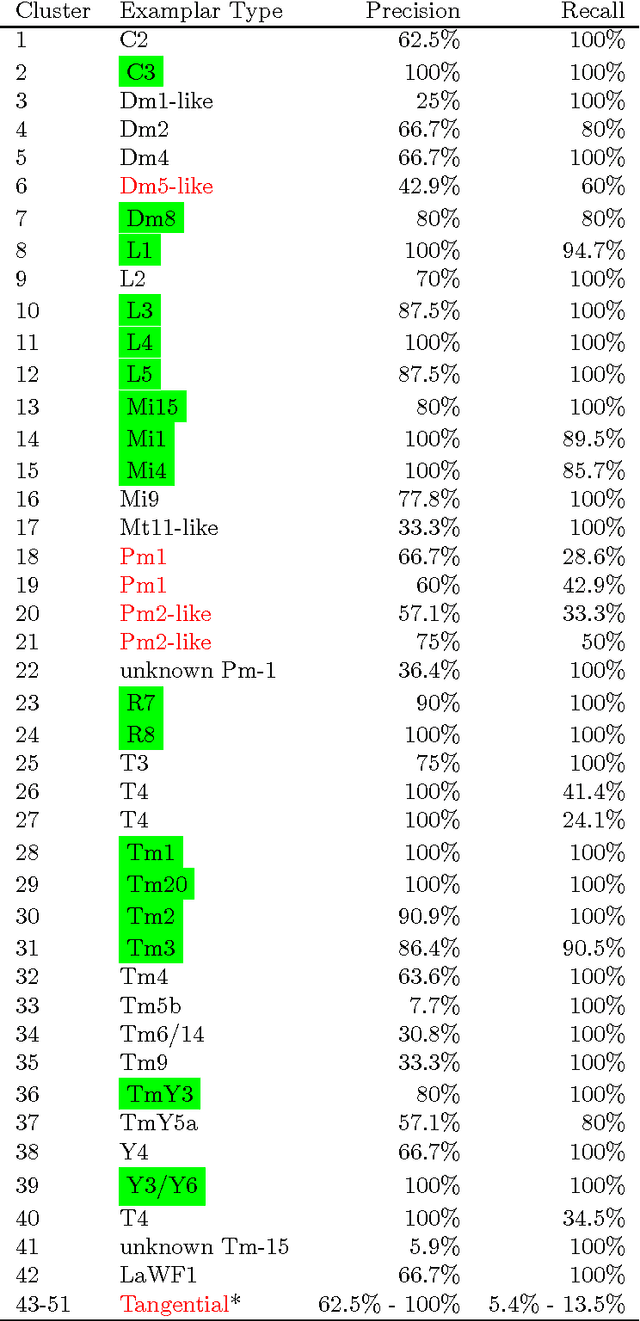
Abstract:Mapping the connectivity of neurons in the brain (i.e., connectomics) is a challenging problem due to both the number of connections in even the smallest organisms and the nanometer resolution required to resolve them. Because of this, previous connectomes contain only hundreds of neurons, such as in the C.elegans connectome. Recent technological advances will unlock the mysteries of increasingly large connectomes (or partial connectomes). However, the value of these maps is limited by our ability to reason with this data and understand any underlying motifs. To aid connectome analysis, we introduce algorithms to cluster similarly-shaped neurons, where 3D neuronal shapes are represented as skeletons. In particular, we propose a novel location-sensitive clustering algorithm. We show clustering results on neurons reconstructed from the Drosophila medulla that show high-accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge