S. Kavitha
Efficient DWT-based fusion techniques using genetic algorithm for optimal parameter estimation
Sep 22, 2020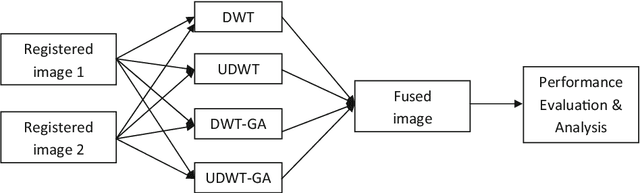

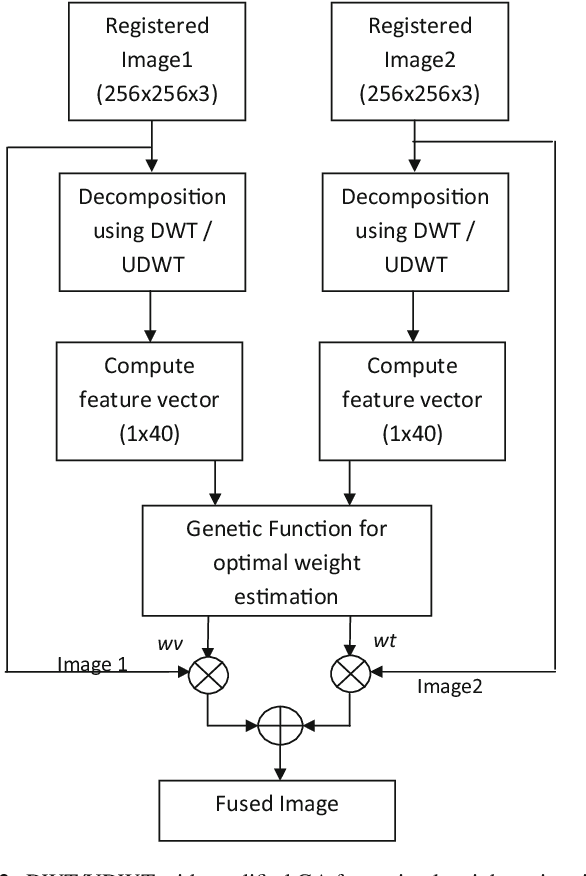

Abstract:Image fusion plays a vital role in medical imaging. Image fusion aims to integrate complementary as well as redundant information from multiple modalities into a single fused image without distortion or loss of information. In this research work, discrete wavelet transform (DWT)and undecimated discrete wavelet transform (UDWT)-based fusion techniques using genetic algorithm (GA)foroptimalparameter(weight)estimationinthefusionprocessareimplemented and analyzed with multi-modality brain images. The lack of shift variance while performing image fusion using DWT is addressed using UDWT. The proposed fusion model uses an efficient, modified GA in DWT and UDWT for optimal parameter estimation, to improve the image quality and contrast. The complexity of the basic GA (pixel level) has been reduced in the modified GA (feature level), by limiting the search space. It is observed from our experiments that fusion using DWT and UDWT techniques with GA for optimal parameter estimation resulted in a better fused image in the aspects of retaining the information and contrast without error, both in human perception as well as evaluation using objective metrics. The contributions of this research work are (1) reduced time and space complexity in estimating the weight values using GA for fusion (2) system is scalable for input image of any size with similar time complexity, owing to feature level GA implementation and (3) identification of source image that contributes more to the fused image, from the weight values estimated.
Features based Mammogram Image Classification using Weighted Feature Support Vector Machine
Sep 19, 2020

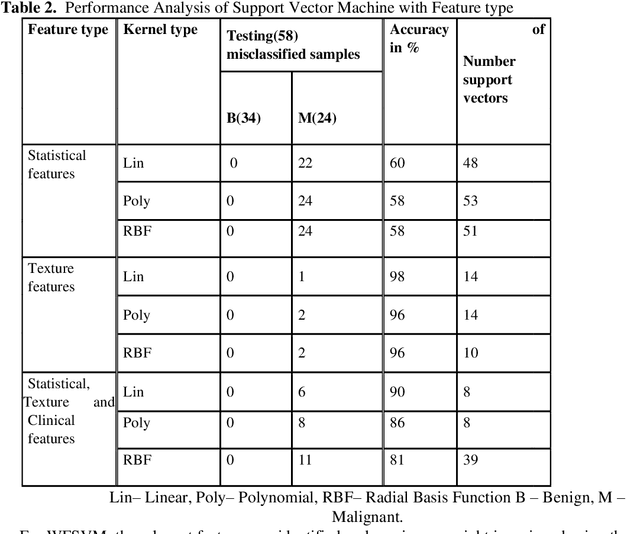

Abstract:In the existing research of mammogram image classification, either clinical data or image features of a specific type is considered along with the supervised classifiers such as Neural Network (NN) and Support Vector Machine (SVM). This paper considers automated classification of breast tissue type as benign or malignant using Weighted Feature Support Vector Machine (WFSVM) through constructing the precomputed kernel function by assigning more weight to relevant features using the principle of maximizing deviations. Initially, MIAS dataset of mammogram images is divided into training and test set, then the preprocessing techniques such as noise removal and background removal are applied to the input images and the Region of Interest (ROI) is identified. The statistical features and texture features are extracted from the ROI and the clinical features are obtained directly from the dataset. The extracted features of the training dataset are used to construct the weighted features and precomputed linear kernel for training the WFSVM, from which the training model file is created. Using this model file the kernel matrix of test samples is classified as benign or malignant. This analysis shows that the texture features have resulted in better accuracy than the other features with WFSVM and SVM. However, the number of support vectors created in WFSVM is less than the SVM classifier.
* 9 pages, 3 figures, "submitted to International Conference on Computing and Communication Systems"
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge