Rene Werner
Anatomy-Informed Deep Learning and Radiomics for Automated Neurofibroma Segmentation in Whole-Body MRI
Feb 21, 2025Abstract:Neurofibromatosis Type 1 is a genetic disorder characterized by the development of neurofibromas (NFs), which exhibit significant variability in size, morphology, and anatomical location. Accurate and automated segmentation of these tumors in whole-body magnetic resonance imaging (WB-MRI) is crucial to assess tumor burden and monitor disease progression. In this study, we present and analyze a fully automated pipeline for NF segmentation in fat-suppressed T2-weighted WB-MRI, consisting of three stages: anatomy segmentation, NF segmentation, and tumor candidate classification. In the first stage, we use the MRSegmentator model to generate an anatomy segmentation mask, extended with a high-risk zone for NFs. This mask is concatenated with the input image as anatomical context information for NF segmentation. The second stage employs an ensemble of 3D anisotropic anatomy-informed U-Nets to produce an NF segmentation confidence mask. In the final stage, tumor candidates are extracted from the confidence mask and classified based on radiomic features, distinguishing tumors from non-tumor regions and reducing false positives. We evaluate the proposed pipeline on three test sets representing different conditions: in-domain data (test set 1), varying imaging protocols and field strength (test set 2), and low tumor burden cases (test set 3). Experimental results show a 68% improvement in per-scan Dice Similarity Coefficient (DSC), a 21% increase in per-tumor DSC, and a two-fold improvement in F1 score for tumor detection in high tumor burden cases by integrating anatomy information. The method is integrated into the 3D Slicer platform for practical clinical use, with the code publicly accessible.
Deep learning-based conditional inpainting for restoration of artifact-affected 4D CT images
Mar 12, 2022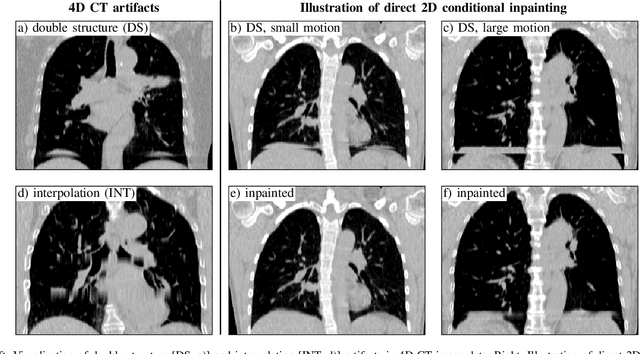
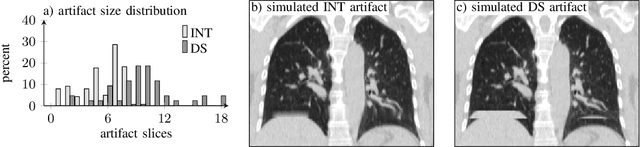
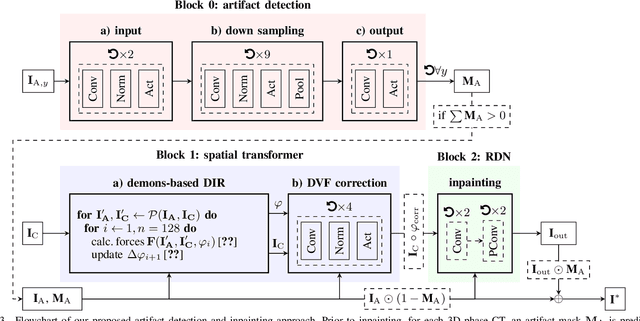
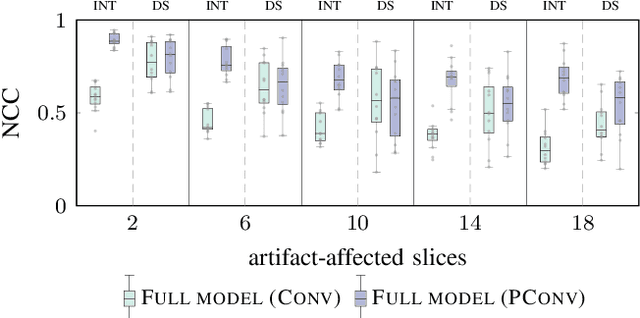
Abstract:4D CT imaging is an essential component of radiotherapy of thoracic/abdominal tumors. 4D CT images are, however, often affected by artifacts that compromise treatment planning quality. In this work, deep learning (DL)-based conditional inpainting is proposed to restore anatomically correct image information of artifact-affected areas. The restoration approach consists of a two-stage process: DL-based detection of common interpolation (INT) and double structure (DS) artifacts, followed by conditional inpainting applied to the artifact areas. In this context, conditional refers to a guidance of the inpainting process by patient-specific image data to ensure anatomically reliable results. Evaluation is based on 65 in-house 4D CT data sets of lung cancer patients (48 with only slight artifacts, 17 with pronounced artifacts) and the publicly available DIRLab 4D CT data (independent external test set). Automated artifact detection revealed a ROC-AUC of 0.99 for INT and 0.97 for DS artifacts (in-house data). The proposed inpainting method decreased the average root mean squared error (RMSE) by 60% (DS) and 42% (INT) for the in-house evaluation data (simulated artifacts for the slight artifact data; original data were considered as ground truth for RMSE computation). For the external DIR-Lab data, the RMSE decreased by 65% and 36%, respectively. Applied to the pronounced artifact data group, on average 68% of the detectable artifacts were removed. The results highlight the potential of DL-based inpainting for the restoration of artifact-affected 4D CT data. Improved performance of conditional inpainting (compared to standard inpainting) illustrates the benefits of exploiting patient-specific prior knowledge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge