Pierre Manceron
RadImageNet-VQA: A Large-Scale CT and MRI Dataset for Radiologic Visual Question Answering
Dec 19, 2025



Abstract:In this work, we introduce RadImageNet-VQA, a large-scale dataset designed to advance radiologic visual question answering (VQA) on CT and MRI exams. Existing medical VQA datasets are limited in scale, dominated by X-ray imaging or biomedical illustrations, and often prone to text-based shortcuts. RadImageNet-VQA is built from expert-curated annotations and provides 750K images paired with 7.5M question-answer samples. It covers three key tasks - abnormality detection, anatomy recognition, and pathology identification - spanning eight anatomical regions and 97 pathology categories, and supports open-ended, closed-ended, and multiple-choice questions. Extensive experiments show that state-of-the-art vision-language models still struggle with fine-grained pathology identification, particularly in open-ended settings and even after fine-tuning. Text-only analysis further reveals that model performance collapses to near-random without image inputs, confirming that RadImageNet-VQA is free from linguistic shortcuts. The full dataset and benchmark are publicly available at https://huggingface.co/datasets/raidium/RadImageNet-VQA.
RAPS-3D: Efficient interactive segmentation for 3D radiological imaging
Jul 10, 2025Abstract:Promptable segmentation, introduced by the Segment Anything Model (SAM), is a promising approach for medical imaging, as it enables clinicians to guide and refine model predictions interactively. However, SAM's architecture is designed for 2D images and does not extend naturally to 3D volumetric data such as CT or MRI scans. Adapting 2D models to 3D typically involves autoregressive strategies, where predictions are propagated slice by slice, resulting in increased inference complexity. Processing large 3D volumes also requires significant computational resources, often leading existing 3D methods to also adopt complex strategies like sliding-window inference to manage memory usage, at the cost of longer inference times and greater implementation complexity. In this paper, we present a simplified 3D promptable segmentation method, inspired by SegVol, designed to reduce inference time and eliminate prompt management complexities associated with sliding windows while achieving state-of-the-art performance.
RadSAM: Segmenting 3D radiological images with a 2D promptable model
Apr 29, 2025Abstract:Medical image segmentation is a crucial and time-consuming task in clinical care, where mask precision is extremely important. The Segment Anything Model (SAM) offers a promising approach, as it provides an interactive interface based on visual prompting and edition to refine an initial segmentation. This model has strong generalization capabilities, does not rely on predefined classes, and adapts to diverse objects; however, it is pre-trained on natural images and lacks the ability to process medical data effectively. In addition, this model is built for 2D images, whereas a whole medical domain is based on 3D images, such as CT and MRI. Recent adaptations of SAM for medical imaging are based on 2D models, thus requiring one prompt per slice to segment 3D objects, making the segmentation process tedious. They also lack important features such as editing. To bridge this gap, we propose RadSAM, a novel method for segmenting 3D objects with a 2D model from a single prompt. In practice, we train a 2D model using noisy masks as initial prompts, in addition to bounding boxes and points. We then use this novel prompt type with an iterative inference pipeline to reconstruct the 3D mask slice-by-slice. We introduce a benchmark to evaluate the model's ability to segment 3D objects in CT images from a single prompt and evaluate the models' out-of-domain transfer and edition capabilities. We demonstrate the effectiveness of our approach against state-of-the-art models on this benchmark using the AMOS abdominal organ segmentation dataset.
ONCOPILOT: A Promptable CT Foundation Model For Solid Tumor Evaluation
Oct 10, 2024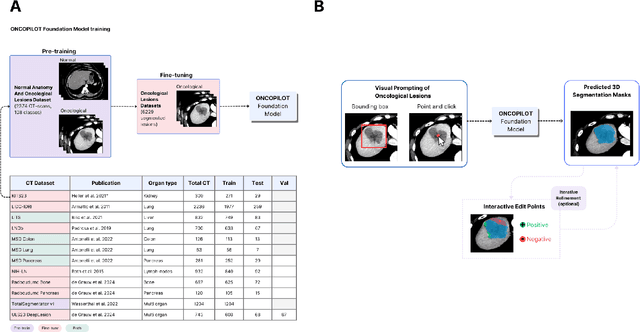
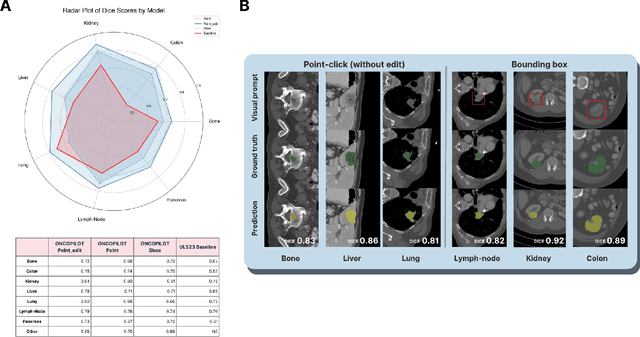
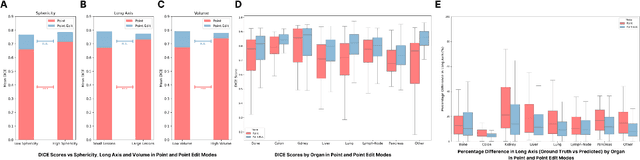
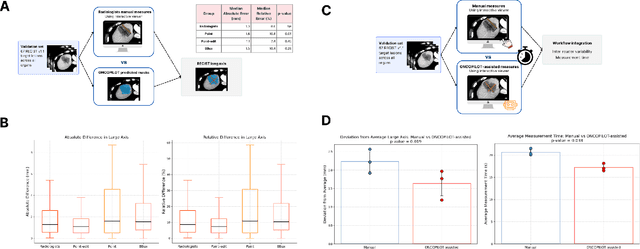
Abstract:Carcinogenesis is a proteiform phenomenon, with tumors emerging in various locations and displaying complex, diverse shapes. At the crucial intersection of research and clinical practice, it demands precise and flexible assessment. However, current biomarkers, such as RECIST 1.1's long and short axis measurements, fall short of capturing this complexity, offering an approximate estimate of tumor burden and a simplistic representation of a more intricate process. Additionally, existing supervised AI models face challenges in addressing the variability in tumor presentations, limiting their clinical utility. These limitations arise from the scarcity of annotations and the models' focus on narrowly defined tasks. To address these challenges, we developed ONCOPILOT, an interactive radiological foundation model trained on approximately 7,500 CT scans covering the whole body, from both normal anatomy and a wide range of oncological cases. ONCOPILOT performs 3D tumor segmentation using visual prompts like point-click and bounding boxes, outperforming state-of-the-art models (e.g., nnUnet) and achieving radiologist-level accuracy in RECIST 1.1 measurements. The key advantage of this foundation model is its ability to surpass state-of-the-art performance while keeping the radiologist in the loop, a capability that previous models could not achieve. When radiologists interactively refine the segmentations, accuracy improves further. ONCOPILOT also accelerates measurement processes and reduces inter-reader variability, facilitating volumetric analysis and unlocking new biomarkers for deeper insights. This AI assistant is expected to enhance the precision of RECIST 1.1 measurements, unlock the potential of volumetric biomarkers, and improve patient stratification and clinical care, while seamlessly integrating into the radiological workflow.
Efficient Medical Question Answering with Knowledge-Augmented Question Generation
May 23, 2024Abstract:In the expanding field of language model applications, medical knowledge representation remains a significant challenge due to the specialized nature of the domain. Large language models, such as GPT-4, obtain reasonable scores on medical question answering tasks, but smaller models are far behind. In this work, we introduce a method to improve the proficiency of a small language model in the medical domain by employing a two-fold approach. We first fine-tune the model on a corpus of medical textbooks. Then, we use GPT-4 to generate questions similar to the downstream task, prompted with textbook knowledge, and use them to fine-tune the model. Additionally, we introduce ECN-QA, a novel medical question answering dataset containing ``progressive questions'' composed of related sequential questions. We show the benefits of our training strategy on this dataset. The study's findings highlight the potential of small language models in the medical domain when appropriately fine-tuned. The code and weights are available at https://github.com/raidium-med/MQG.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge