Nazli Tümer
Shape Modeling of Longitudinal Medical Images: From Diffeomorphic Metric Mapping to Deep Learning
Mar 27, 2025Abstract:Living biological tissue is a complex system, constantly growing and changing in response to external and internal stimuli. These processes lead to remarkable and intricate changes in shape. Modeling and understanding both natural and pathological (or abnormal) changes in the shape of anatomical structures is highly relevant, with applications in diagnostic, prognostic, and therapeutic healthcare. Nevertheless, modeling the longitudinal shape change of biological tissue is a non-trivial task due to its inherent nonlinear nature. In this review, we highlight several existing methodologies and tools for modeling longitudinal shape change (i.e., spatiotemporal shape modeling). These methods range from diffeomorphic metric mapping to deep-learning based approaches (e.g., autoencoders, generative networks, recurrent neural networks, etc.). We discuss the synergistic combinations of existing technologies and potential directions for future research, underscoring key deficiencies in the current research landscape.
Early detection of knee osteoarthritis using deep learning on knee magnetic resonance images
Sep 02, 2022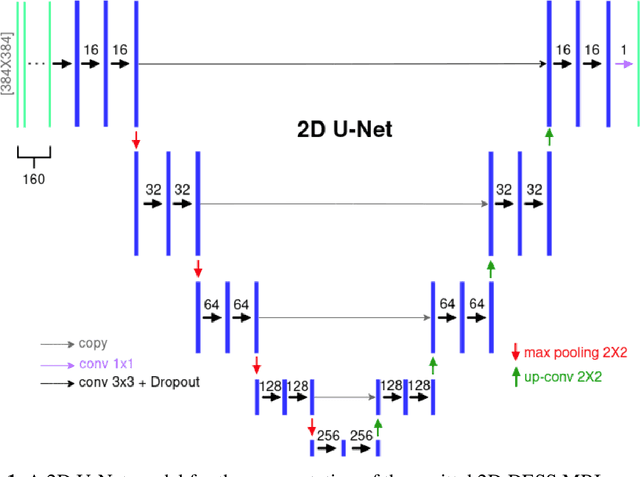

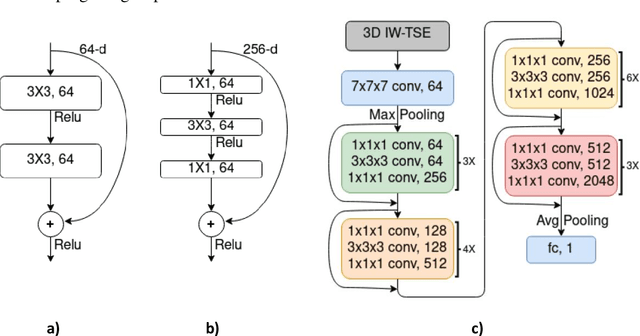
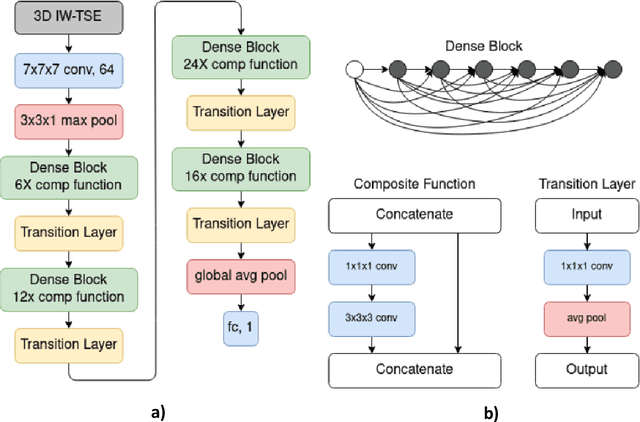
Abstract:The aim of this study was to investigate the influence of MRI and patient data on the prediction of knee osteoarthritis (OA) incidence using different deep learning architectures. Knee OA incidence within 24 months was predicted using the intermediate-weighted turbo spin-echo (IW-TSE) sequence of 593 patients from the Osteoarthritis Initiative. To extract a region of interest containing the knee joint from the IW-TSE sequence, a U-Net model was trained and used to segment bone on a dual echo steady state (DESS) sequence. Subsequently, IW-TSE and DESS sequences were registered and the DESS segmentations were transformed to the corresponding IW-TSE scans. The performance of MRI-based features in the prediction of knee OA incidence was tested using three different deep learning architectures: a residual network (ResNet), a densely connected convolutional network (DenseNet), and a convolutional variational autoencoder (CVAE). To evaluate the predictive performance of MRI-based features alone, the outputs of ResNet, DenseNet, and CVAE were coupled with patient data (i.e., age, gender, BMI) and used as input to a Logistic Regression (LR) Classifier. Knee OA was defined based on visual MRI and X-ray-based OA features. The ResNet and DenseNet showed similar results, with both methods having the area under the receiver operating characteristic curve (AUC) values up to 0.6269. The best performing OA detection model was CVAE with an AUC of 0.6699 when combined with patient data and an AUC of 0.6689 when used alone as input to the LR classifier. The results showed that three deep learning algorithms have similar metrics when using IW-TSE MRIs and their performance increased with the inclusion of patient data, which shows the strong influence of variables such as age, gender, and BMI on the detection of knee OA.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge