Lennart P. L. Landsmeer
Spatial Spiking Neural Networks Enable Efficient and Robust Temporal Computation
Dec 17, 2025Abstract:The efficiency of modern machine intelligence depends on high accuracy with minimal computational cost. In spiking neural networks (SNNs), synaptic delays are crucial for encoding temporal structure, yet existing models treat them as fully trainable, unconstrained parameters, leading to large memory footprints, higher computational demand, and a departure from biological plausibility. In the brain, however, delays arise from physical distances between neurons embedded in space. Building on this principle, we introduce Spatial Spiking Neural Networks (SpSNNs), a framework in which neurons learn coordinates in a finite-dimensional Euclidean space and delays emerge from inter-neuron distances. This replaces per-synapse delay learning with position learning, substantially reducing parameter count while retaining temporal expressiveness. Across the Yin-Yang and Spiking Heidelberg Digits benchmarks, SpSNNs outperform SNNs with unconstrained delays despite using far fewer parameters. Performance consistently peaks in 2D and 3D networks rather than infinite-dimensional delay spaces, revealing a geometric regularization effect. Moreover, dynamically sparsified SpSNNs maintain full accuracy even at 90% sparsity, matching standard delay-trained SNNs while using up to 18x fewer parameters. Because learned spatial layouts map naturally onto hardware geometries, SpSNNs lend themselves to efficient neuromorphic implementation. Methodologically, SpSNNs compute exact delay gradients via automatic differentiation with custom-derived rules, supporting arbitrary neuron models and architectures. Altogether, SpSNNs provide a principled platform for exploring spatial structure in temporal computation and offer a hardware-friendly substrate for scalable, energy-efficient neuromorphic intelligence.
A Lightweight Architecture for Real-Time Neuronal-Spike Classification
Nov 08, 2023Abstract:Electrophysiological recordings of neural activity in a mouse's brain are very popular among neuroscientists for understanding brain function. One particular area of interest is acquiring recordings from the Purkinje cells in the cerebellum in order to understand brain injuries and the loss of motor functions. However, current setups for such experiments do not allow the mouse to move freely and, thus, do not capture its natural behaviour since they have a wired connection between the animal's head stage and an acquisition device. In this work, we propose a lightweight neuronal-spike detection and classification architecture that leverages on the unique characteristics of the Purkinje cells to discard unneeded information from the sparse neural data in real time. This allows the (condensed) data to be easily stored on a removable storage device on the head stage, alleviating the need for wires. Our proposed implementation shows a >95% overall classification accuracy while still resulting in a small-form-factor design, which allows for the free movement of mice during experiments. Moreover, the power-efficient nature of the design and the usage of STT-RAM (Spin Transfer Torque Magnetic Random Access Memory) as the removable storage allows the head stage to easily operate on a tiny battery for up to approximately 4 days.
Tricking AI chips into Simulating the Human Brain: A Detailed Performance Analysis
Jan 31, 2023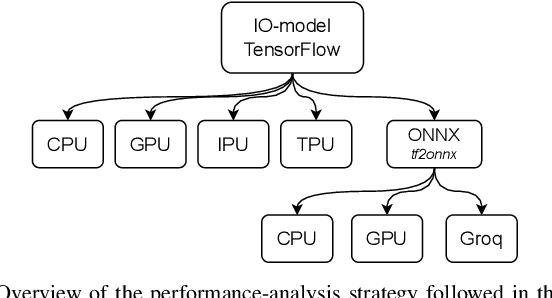
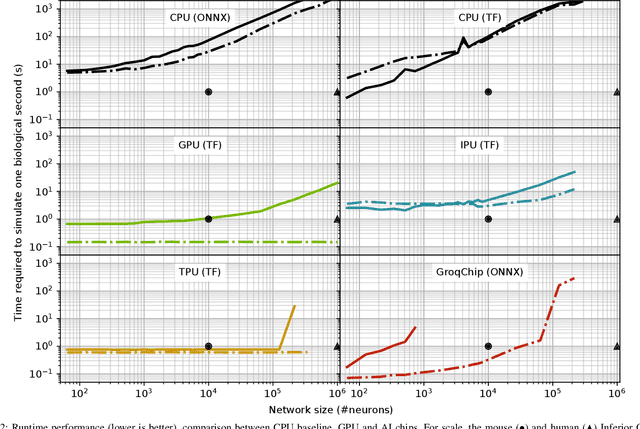
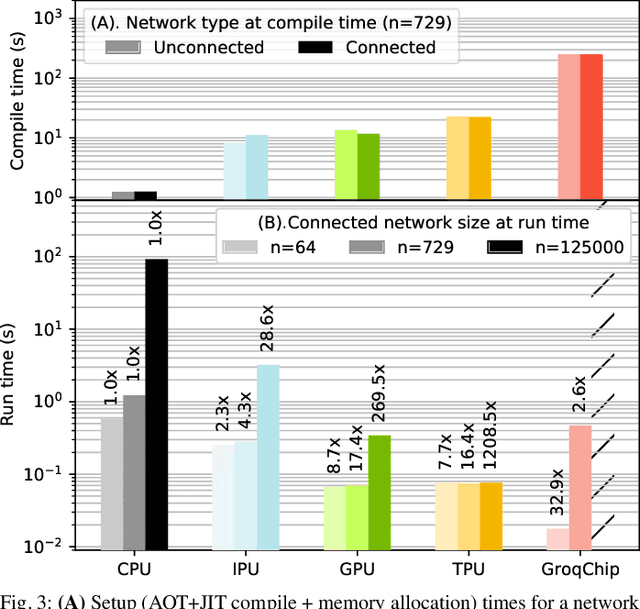
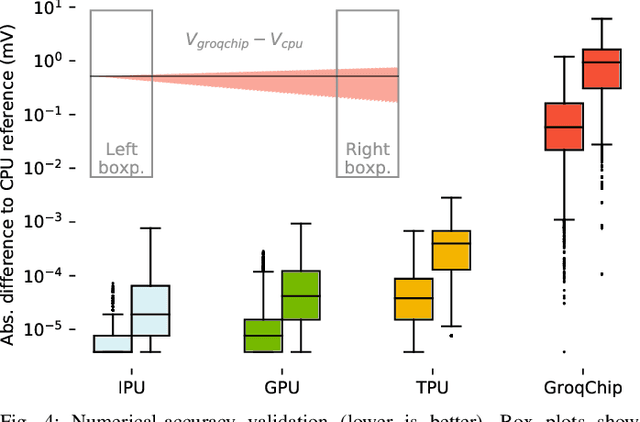
Abstract:Challenging the Nvidia monopoly, dedicated AI-accelerator chips have begun emerging for tackling the computational challenge that the inference and, especially, the training of modern deep neural networks (DNNs) poses to modern computers. The field has been ridden with studies assessing the performance of these contestants across various DNN model types. However, AI-experts are aware of the limitations of current DNNs and have been working towards the fourth AI wave which will, arguably, rely on more biologically inspired models, predominantly on spiking neural networks (SNNs). At the same time, GPUs have been heavily used for simulating such models in the field of computational neuroscience, yet AI-chips have not been tested on such workloads. The current paper aims at filling this important gap by evaluating multiple, cutting-edge AI-chips (Graphcore IPU, GroqChip, Nvidia GPU with Tensor Cores and Google TPU) on simulating a highly biologically detailed model of a brain region, the inferior olive (IO). This IO application stress-tests the different AI-platforms for highlighting architectural tradeoffs by varying its compute density, memory requirements and floating-point numerical accuracy. Our performance analysis reveals that the simulation problem maps extremely well onto the GPU and TPU architectures, which for networks of 125,000 cells leads to a 28x respectively 1,208x speedup over CPU runtimes. At this speed, the TPU sets a new record for largest real-time IO simulation. The GroqChip outperforms both platforms for small networks but, due to implementing some floating-point operations at reduced accuracy, is found not yet usable for brain simulation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge