Karthik Soman
Weill Institute for Neurosciences, Department of Neurology, University of California San Francisco, San Francisco, CA, USA
Zebra-Llama: A Context-Aware Large Language Model for Democratizing Rare Disease Knowledge
Nov 04, 2024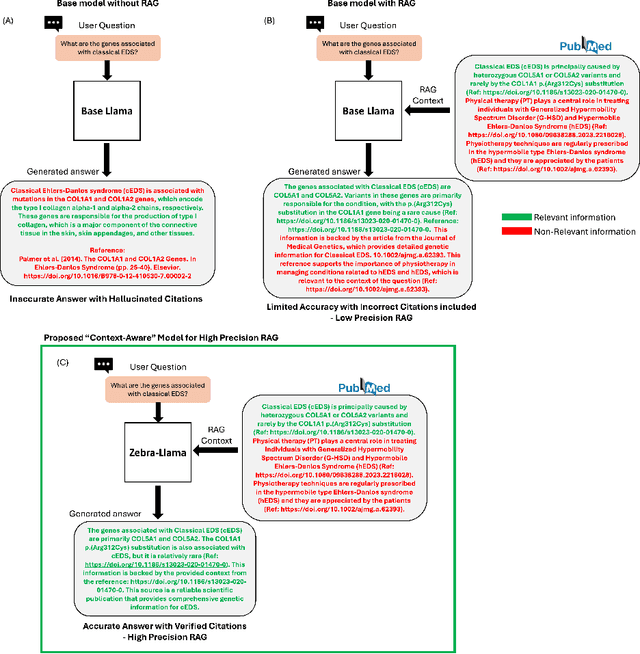
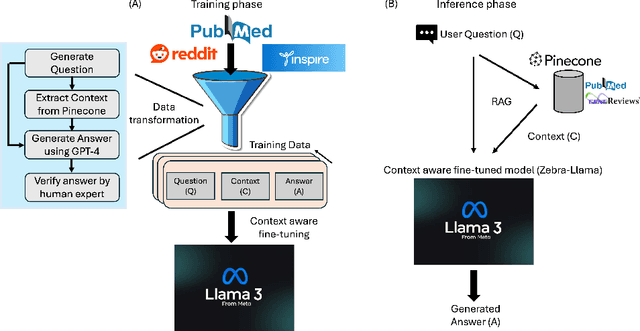
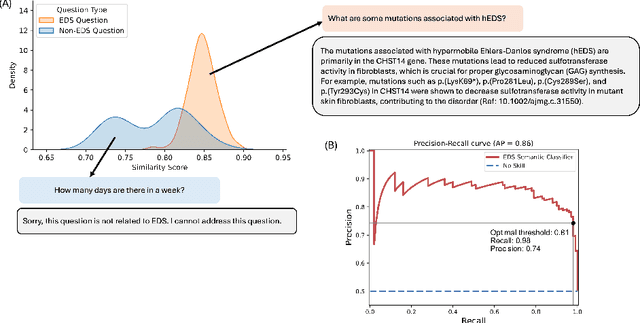
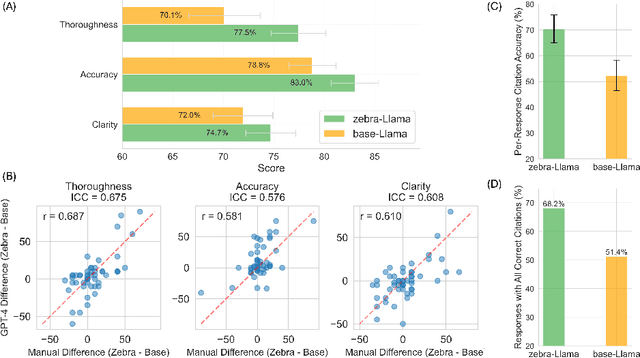
Abstract:Rare diseases present unique challenges in healthcare, often suffering from delayed diagnosis and fragmented information landscapes. The scarcity of reliable knowledge in these conditions poses a distinct challenge for Large Language Models (LLMs) in supporting clinical management and delivering precise patient information underscoring the need for focused training on these 'zebra' cases. We present Zebra-Llama, a specialized context-aware language model with high precision Retrieval Augmented Generation (RAG) capability, focusing on Ehlers-Danlos Syndrome (EDS) as our case study. EDS, affecting 1 in 5,000 individuals, exemplifies the complexities of rare diseases with its diverse symptoms, multiple subtypes, and evolving diagnostic criteria. By implementing a novel context-aware fine-tuning methodology trained on questions derived from medical literature, patient experiences, and clinical resources, along with expertly curated responses, Zebra-Llama demonstrates unprecedented capabilities in handling EDS-related queries. On a test set of real-world questions collected from EDS patients and clinicians, medical experts evaluated the responses generated by both models, revealing Zebra-Llama's substantial improvements over base model (Llama 3.1-8B-Instruct) in thoroughness (77.5% vs. 70.1%), accuracy (83.0% vs. 78.8%), clarity (74.7% vs. 72.0%) and citation reliability (70.6% vs. 52.3%). Released as an open-source resource, Zebra-Llama not only provides more accessible and reliable EDS information but also establishes a framework for developing specialized AI solutions for other rare conditions. This work represents a crucial step towards democratizing expert-level knowledge in rare disease management, potentially transforming how healthcare providers and patients navigate the complex landscape of rare diseases.
Biomedical knowledge graph-enhanced prompt generation for large language models
Nov 29, 2023



Abstract:Large Language Models (LLMs) have been driving progress in AI at an unprecedented rate, yet still face challenges in knowledge-intensive domains like biomedicine. Solutions such as pre-training and domain-specific fine-tuning add substantial computational overhead, and the latter require domain-expertise. External knowledge infusion is task-specific and requires model training. Here, we introduce a task-agnostic Knowledge Graph-based Retrieval Augmented Generation (KG-RAG) framework by leveraging the massive biomedical KG SPOKE with LLMs such as Llama-2-13b, GPT-3.5-Turbo and GPT-4, to generate meaningful biomedical text rooted in established knowledge. KG-RAG consistently enhanced the performance of LLMs across various prompt types, including one-hop and two-hop prompts, drug repurposing queries, biomedical true/false questions, and multiple-choice questions (MCQ). Notably, KG-RAG provides a remarkable 71% boost in the performance of the Llama-2 model on the challenging MCQ dataset, demonstrating the framework's capacity to empower open-source models with fewer parameters for domain-specific questions. Furthermore, KG-RAG enhanced the performance of proprietary GPT models, such as GPT-3.5 which exhibited improvement over GPT-4 in context utilization on MCQ data. Our approach was also able to address drug repurposing questions, returning meaningful repurposing suggestions. In summary, the proposed framework combines explicit and implicit knowledge of KG and LLM, respectively, in an optimized fashion, thus enhancing the adaptability of general-purpose LLMs to tackle domain-specific questions in a unified framework.
Beyond Low Earth Orbit: Biomonitoring, Artificial Intelligence, and Precision Space Health
Dec 22, 2021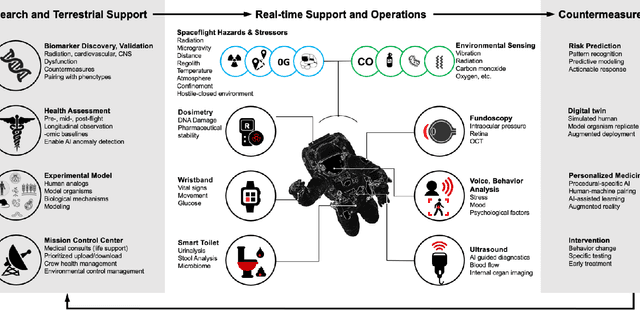
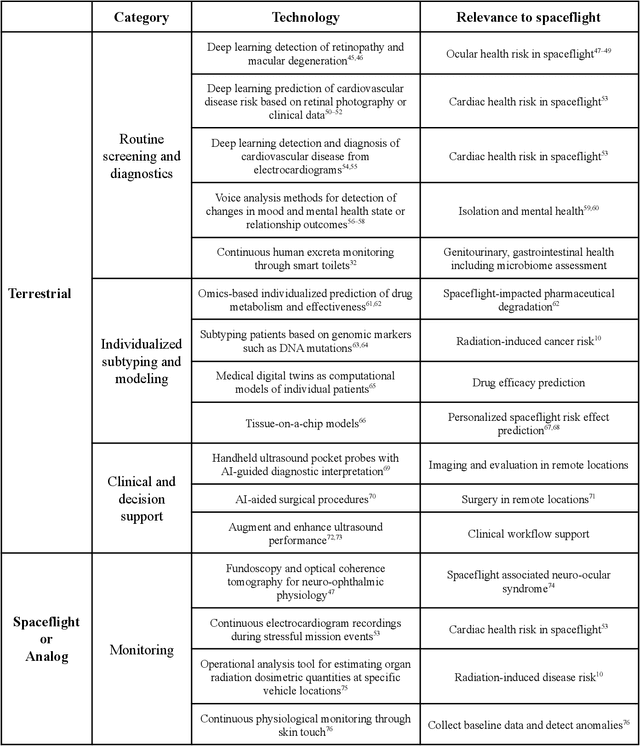
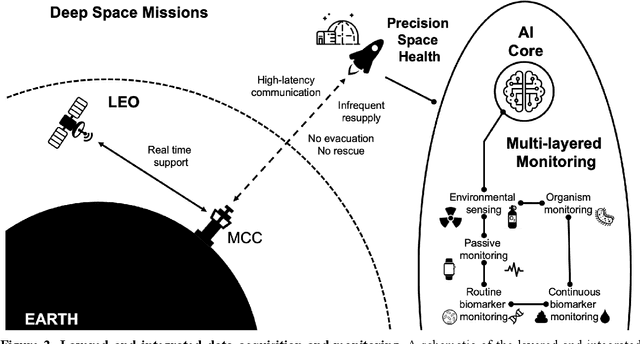
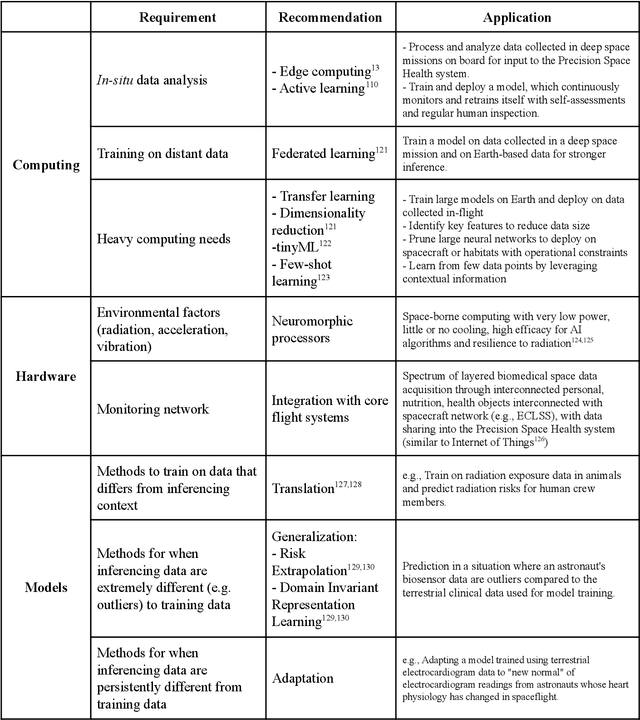
Abstract:Human space exploration beyond low Earth orbit will involve missions of significant distance and duration. To effectively mitigate myriad space health hazards, paradigm shifts in data and space health systems are necessary to enable Earth-independence, rather than Earth-reliance. Promising developments in the fields of artificial intelligence and machine learning for biology and health can address these needs. We propose an appropriately autonomous and intelligent Precision Space Health system that will monitor, aggregate, and assess biomedical statuses; analyze and predict personalized adverse health outcomes; adapt and respond to newly accumulated data; and provide preventive, actionable, and timely insights to individual deep space crew members and iterative decision support to their crew medical officer. Here we present a summary of recommendations from a workshop organized by the National Aeronautics and Space Administration, on future applications of artificial intelligence in space biology and health. In the next decade, biomonitoring technology, biomarker science, spacecraft hardware, intelligent software, and streamlined data management must mature and be woven together into a Precision Space Health system to enable humanity to thrive in deep space.
Beyond Low Earth Orbit: Biological Research, Artificial Intelligence, and Self-Driving Labs
Dec 22, 2021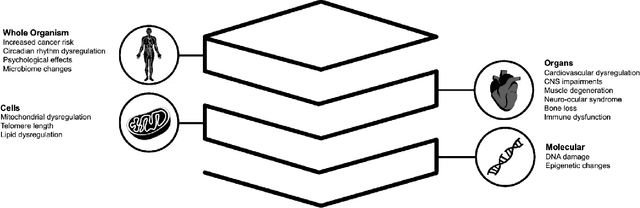
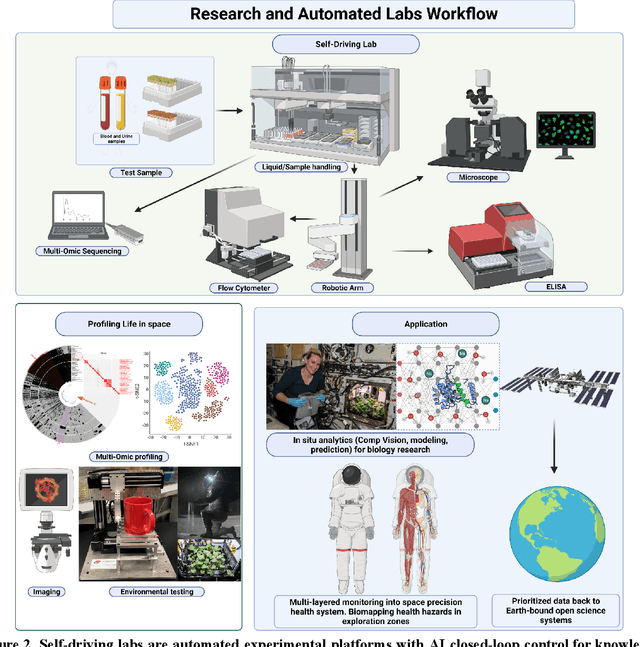
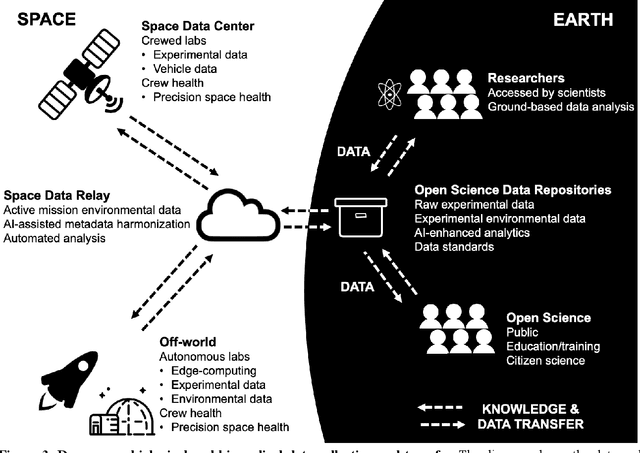
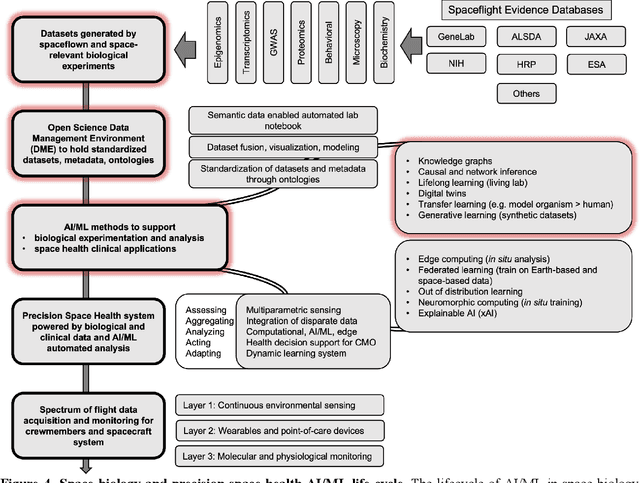
Abstract:Space biology research aims to understand fundamental effects of spaceflight on organisms, develop foundational knowledge to support deep space exploration, and ultimately bioengineer spacecraft and habitats to stabilize the ecosystem of plants, crops, microbes, animals, and humans for sustained multi-planetary life. To advance these aims, the field leverages experiments, platforms, data, and model organisms from both spaceborne and ground-analog studies. As research is extended beyond low Earth orbit, experiments and platforms must be maximally autonomous, light, agile, and intelligent to expedite knowledge discovery. Here we present a summary of recommendations from a workshop organized by the National Aeronautics and Space Administration on artificial intelligence, machine learning, and modeling applications which offer key solutions toward these space biology challenges. In the next decade, the synthesis of artificial intelligence into the field of space biology will deepen the biological understanding of spaceflight effects, facilitate predictive modeling and analytics, support maximally autonomous and reproducible experiments, and efficiently manage spaceborne data and metadata, all with the goal to enable life to thrive in deep space.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge