Gordon Harris
LNQ 2023 challenge: Benchmark of weakly-supervised techniques for mediastinal lymph node quantification
Aug 19, 2024



Abstract:Accurate assessment of lymph node size in 3D CT scans is crucial for cancer staging, therapeutic management, and monitoring treatment response. Existing state-of-the-art segmentation frameworks in medical imaging often rely on fully annotated datasets. However, for lymph node segmentation, these datasets are typically small due to the extensive time and expertise required to annotate the numerous lymph nodes in 3D CT scans. Weakly-supervised learning, which leverages incomplete or noisy annotations, has recently gained interest in the medical imaging community as a potential solution. Despite the variety of weakly-supervised techniques proposed, most have been validated only on private datasets or small publicly available datasets. To address this limitation, the Mediastinal Lymph Node Quantification (LNQ) challenge was organized in conjunction with the 26th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2023). This challenge aimed to advance weakly-supervised segmentation methods by providing a new, partially annotated dataset and a robust evaluation framework. A total of 16 teams from 5 countries submitted predictions to the validation leaderboard, and 6 teams from 3 countries participated in the evaluation phase. The results highlighted both the potential and the current limitations of weakly-supervised approaches. On one hand, weakly-supervised approaches obtained relatively good performance with a median Dice score of $61.0\%$. On the other hand, top-ranked teams, with a median Dice score exceeding $70\%$, boosted their performance by leveraging smaller but fully annotated datasets to combine weak supervision and full supervision. This highlights both the promise of weakly-supervised methods and the ongoing need for high-quality, fully annotated data to achieve higher segmentation performance.
DINs: Deep Interactive Networks for Neurofibroma Segmentation in Neurofibromatosis Type 1 on Whole-Body MRI
Jun 07, 2021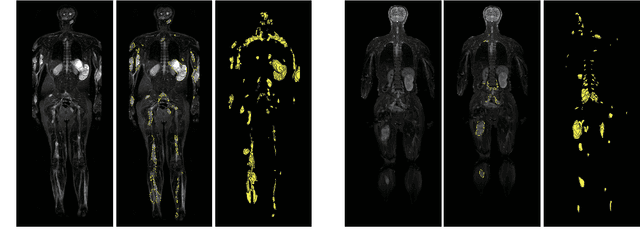
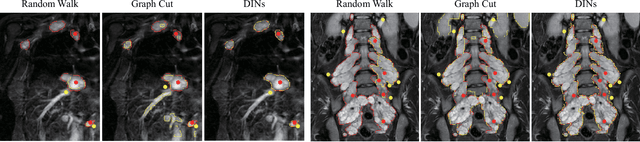
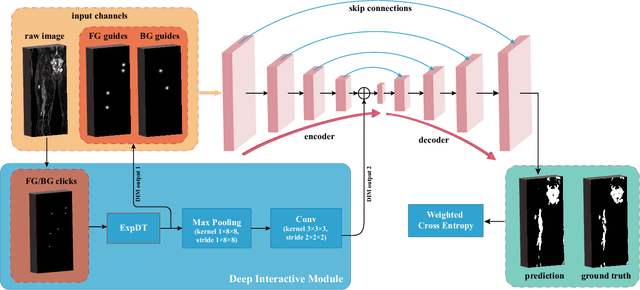
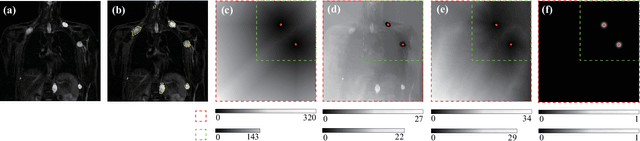
Abstract:Neurofibromatosis type 1 (NF1) is an autosomal dominant tumor predisposition syndrome that involves the central and peripheral nervous systems. Accurate detection and segmentation of neurofibromas are essential for assessing tumor burden and longitudinal tumor size changes. Automatic convolutional neural networks (CNNs) are sensitive and vulnerable as tumors' variable anatomical location and heterogeneous appearance on MRI. In this study, we propose deep interactive networks (DINs) to address the above limitations. User interactions guide the model to recognize complicated tumors and quickly adapt to heterogeneous tumors. We introduce a simple but effective Exponential Distance Transform (ExpDT) that converts user interactions into guide maps regarded as the spatial and appearance prior. Comparing with popular Euclidean and geodesic distances, ExpDT is more robust to various image sizes, which reserves the distribution of interactive inputs. Furthermore, to enhance the tumor-related features, we design a deep interactive module to propagate the guides into deeper layers. We train and evaluate DINs on three MRI data sets from NF1 patients. The experiment results yield significant improvements of 44% and 14% in DSC comparing with automated and other interactive methods, respectively. We also experimentally demonstrate the efficiency of DINs in reducing user burden when comparing with conventional interactive methods. The source code of our method is available at \url{https://github.com/Jarvis73/DINs}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge