Daniel Parra
Learning Difference Equations with Structured Grammatical Evolution for Postprandial Glycaemia Prediction
Jul 03, 2023



Abstract:People with diabetes must carefully monitor their blood glucose levels, especially after eating. Blood glucose regulation requires a proper combination of food intake and insulin boluses. Glucose prediction is vital to avoid dangerous post-meal complications in treating individuals with diabetes. Although traditional methods, such as artificial neural networks, have shown high accuracy rates, sometimes they are not suitable for developing personalised treatments by physicians due to their lack of interpretability. In this study, we propose a novel glucose prediction method emphasising interpretability: Interpretable Sparse Identification by Grammatical Evolution. Combined with a previous clustering stage, our approach provides finite difference equations to predict postprandial glucose levels up to two hours after meals. We divide the dataset into four-hour segments and perform clustering based on blood glucose values for the twohour window before the meal. Prediction models are trained for each cluster for the two-hour windows after meals, allowing predictions in 15-minute steps, yielding up to eight predictions at different time horizons. Prediction safety was evaluated based on Parkes Error Grid regions. Our technique produces safe predictions through explainable expressions, avoiding zones D (0.2% average) and E (0%) and reducing predictions on zone C (6.2%). In addition, our proposal has slightly better accuracy than other techniques, including sparse identification of non-linear dynamics and artificial neural networks. The results demonstrate that our proposal provides interpretable solutions without sacrificing prediction accuracy, offering a promising approach to glucose prediction in diabetes management that balances accuracy, interpretability, and computational efficiency.
Identifying Differential Equations to predict Blood Glucose using Sparse Identification of Nonlinear Systems
Sep 28, 2022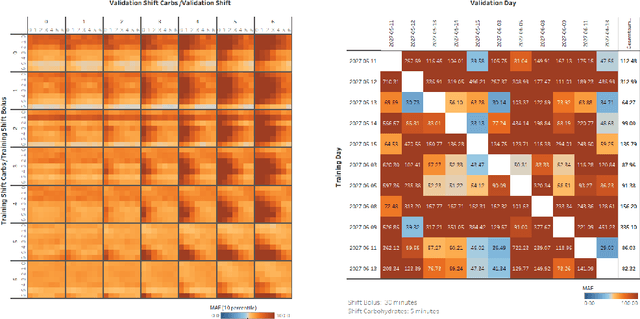
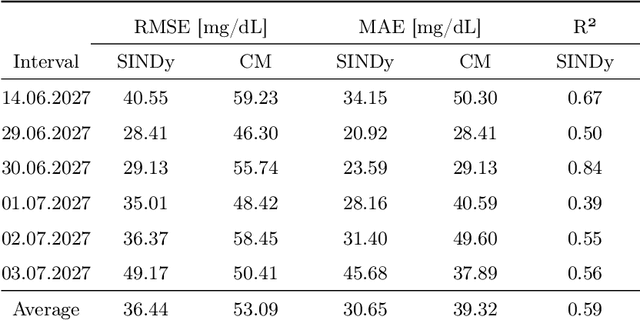

Abstract:Describing dynamic medical systems using machine learning is a challenging topic with a wide range of applications. In this work, the possibility of modeling the blood glucose level of diabetic patients purely on the basis of measured data is described. A combination of the influencing variables insulin and calories are used to find an interpretable model. The absorption speed of external substances in the human body depends strongly on external influences, which is why time-shifts are added for the influencing variables. The focus is put on identifying the best timeshifts that provide robust models with good prediction accuracy that are independent of other unknown external influences. The modeling is based purely on the measured data using Sparse Identification of Nonlinear Dynamics. A differential equation is determined which, starting from an initial value, simulates blood glucose dynamics. By applying the best model to test data, we can show that it is possible to simulate the long-term blood glucose dynamics using differential equations and few, influencing variables.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge