Basma Nasir
TagGAN: A Generative Model for Data Tagging
Feb 25, 2025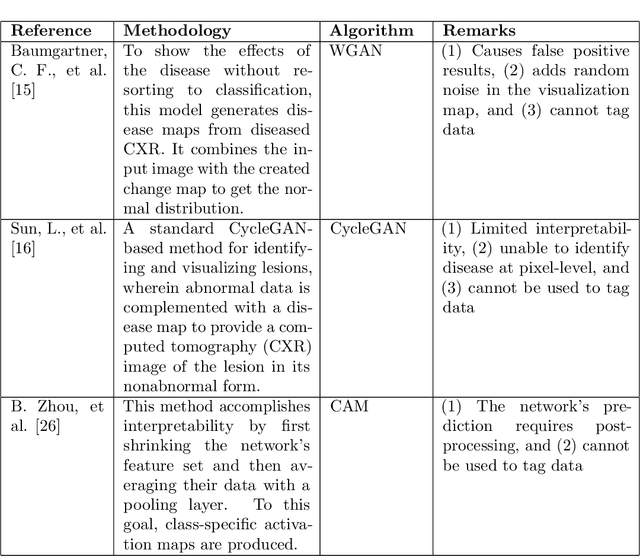
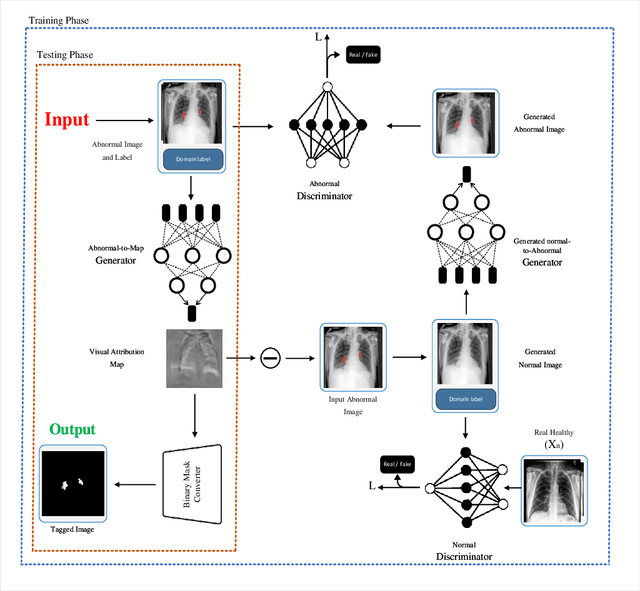
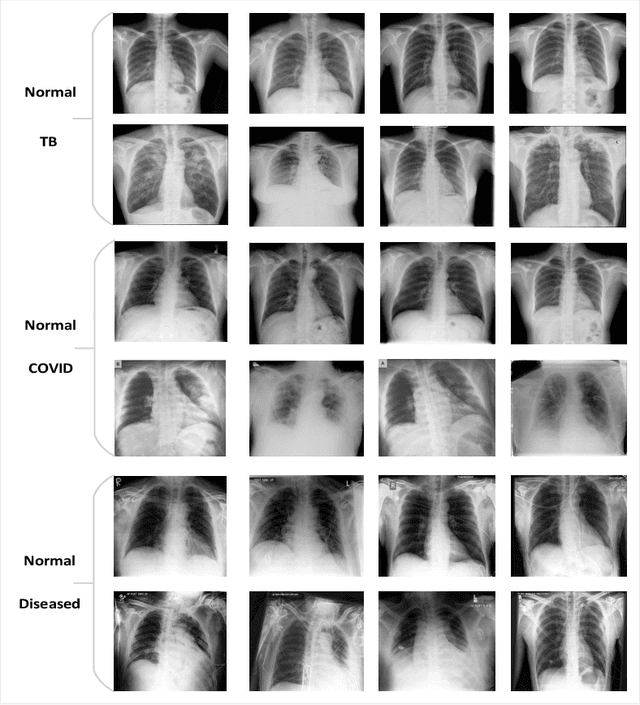

Abstract:Precise identification and localization of disease-specific features at the pixel-level are particularly important for early diagnosis, disease progression monitoring, and effective treatment in medical image analysis. However, conventional diagnostic AI systems lack decision transparency and cannot operate well in environments where there is a lack of pixel-level annotations. In this study, we propose a novel Generative Adversarial Networks (GANs)-based framework, TagGAN, which is tailored for weakly-supervised fine-grained disease map generation from purely image-level labeled data. TagGAN generates a pixel-level disease map during domain translation from an abnormal image to a normal representation. Later, this map is subtracted from the input abnormal image to convert it into its normal counterpart while preserving all the critical anatomical details. Our method is first to generate fine-grained disease maps to visualize disease lesions in a weekly supervised setting without requiring pixel-level annotations. This development enhances the interpretability of diagnostic AI by providing precise visualizations of disease-specific regions. It also introduces automated binary mask generation to assist radiologists. Empirical evaluations carried out on the benchmark datasets, CheXpert, TBX11K, and COVID-19, demonstrate the capability of TagGAN to outperform current top models in accurately identifying disease-specific pixels. This outcome highlights the capability of the proposed model to tag medical images, significantly reducing the workload for radiologists by eliminating the need for binary masks during training.
Weakly Supervised Pixel-Level Annotation with Visual Interpretability
Feb 25, 2025



Abstract:Medical image annotation is essential for diagnosing diseases, yet manual annotation is time-consuming, costly, and prone to variability among experts. To address these challenges, we propose an automated explainable annotation system that integrates ensemble learning, visual explainability, and uncertainty quantification. Our approach combines three pre-trained deep learning models - ResNet50, EfficientNet, and DenseNet - enhanced with XGrad-CAM for visual explanations and Monte Carlo Dropout for uncertainty quantification. This ensemble mimics the consensus of multiple radiologists by intersecting saliency maps from models that agree on the diagnosis while uncertain predictions are flagged for human review. We evaluated our system using the TBX11K medical imaging dataset and a Fire segmentation dataset, demonstrating its robustness across different domains. Experimental results show that our method outperforms baseline models, achieving 93.04% accuracy on TBX11K and 96.4% accuracy on the Fire dataset. Moreover, our model produces precise pixel-level annotations despite being trained with only image-level labels, achieving Intersection over Union IoU scores of 36.07% and 64.7%, respectively. By enhancing the accuracy and interpretability of image annotations, our approach offers a reliable and transparent solution for medical diagnostics and other image analysis tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge