Abicumaran Uthamacumaran
Machine Intelligence-Driven Classification of Cancer Patients-Derived Extracellular Vesicles using Fluorescence Correlation Spectroscopy: Results from a Pilot Study
Feb 01, 2022Abstract:Patient-derived extracellular vesicles (EVs) that contains a complex biological cargo is a valuable source of liquid biopsy diagnostics to aid in early detection, cancer screening, and precision nanotherapeutics. In this study, we predicted that coupling cancer patient blood-derived EVs to time-resolved spectroscopy and artificial intelligence (AI) could provide a robust cancer screening and follow-up tools. Methods: Fluorescence correlation spectroscopy (FCS) measurements were performed on 24 blood samples-derived EVs. Blood samples were obtained from 15 cancer patients (presenting 5 different types of cancers), and 9 healthy controls (including patients with benign lesions). The obtained FCS autocorrelation spectra were processed into power spectra using the Fast-Fourier Transform algorithm and subjected to various machine learning algorithms to distinguish cancer spectra from healthy control spectra. Results and Applications: The performance of AdaBoost Random Forest (RF) classifier, support vector machine, and multilayer perceptron, were tested on selected frequencies in the N=118 power spectra. The RF classifier exhibited a 90% classification accuracy and high sensitivity and specificity in distinguishing the FCS power spectra of cancer patients from those of healthy controls. Further, an image convolutional neural network (CNN), ResNet network, and a quantum CNN were assessed on the power spectral images as additional validation tools. All image-based CNNs exhibited a nearly equal classification performance with an accuracy of roughly 82% and reasonably high sensitivity and specificity scores. Our pilot study demonstrates that AI-algorithms coupled to time-resolved FCS power spectra can accurately and differentially classify the complex patient-derived EVs from different cancer samples of distinct tissue subtypes.
Machine Learning Characterization of Cancer Patients-Derived Extracellular Vesicles using Vibrational Spectroscopies
Aug 25, 2021
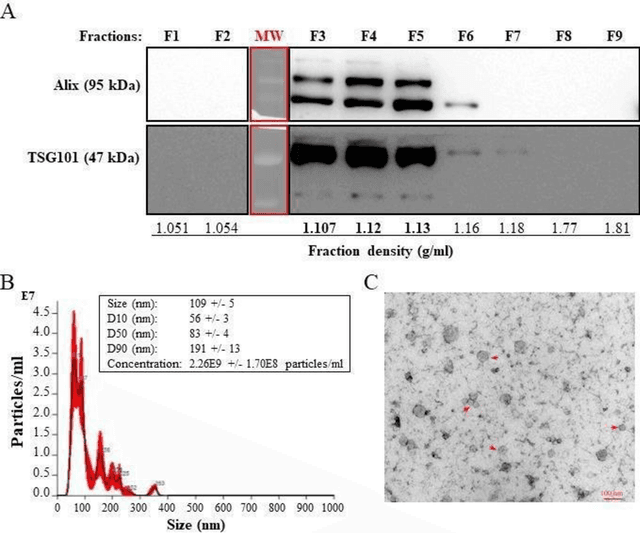

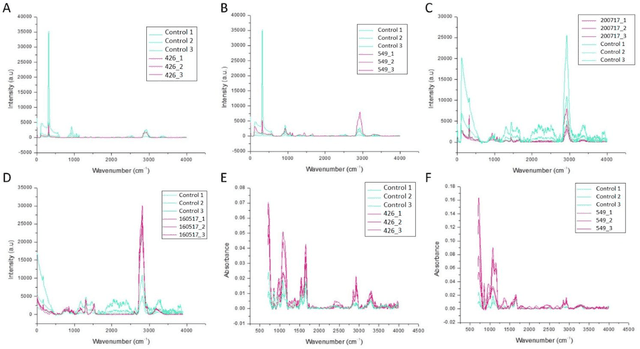
Abstract:The early detection of cancer is a challenging problem in medicine. The blood sera of cancer patients are enriched with heterogeneous secretory lipid bound extracellular vesicles (EVs), which present a complex repertoire of information and biomarkers, representing their cell of origin, that are being currently studied in the field of liquid biopsy and cancer screening. Vibrational spectroscopies provide non-invasive approaches for the assessment of structural and biophysical properties in complex biological samples. In this study, multiple Raman spectroscopy measurements were performed on the EVs extracted from the blood sera of 9 patients consisting of four different cancer subtypes (colorectal cancer, hepatocellular carcinoma, breast cancer and pancreatic cancer) and five healthy patients (controls). FTIR(Fourier Transform Infrared) spectroscopy measurements were performed as a complementary approach to Raman analysis, on two of the four cancer subtypes. The AdaBoost Random Forest Classifier, Decision Trees, and Support Vector Machines (SVM) distinguished the baseline corrected Raman spectra of cancer EVs from those of healthy controls (18 spectra) with a classification accuracy of greater than 90% when reduced to a spectral frequency range of 1800 to 1940 inverse cm, and subjected to a 0.5 training/testing split. FTIR classification accuracy on 14 spectra showed an 80% classification accuracy. Our findings demonstrate that basic machine learning algorithms are powerful tools to distinguish the complex vibrational spectra of cancer patient EVs from those of healthy patients. These experimental methods hold promise as valid and efficient liquid biopsy for machine intelligence-assisted early cancer screening.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge