Transform-Domain Classification of Human Cells based on DNA Methylation Datasets
Paper and Code
Dec 31, 2019
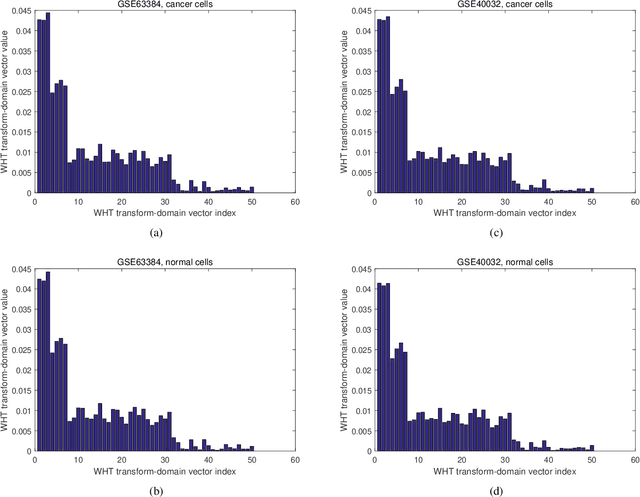


A novel method to classify human cells is presented in this work based on the transform-domain method on DNA methylation data. DNA methylation profile variations are observed in human cells with the progression of disease stages, and the proposal is based on this DNA methylation variation to classify normal and disease cells including cancer cells. The cancer cell types investigated in this work cover hepatocellular (sample size n = 40), colorectal (n = 44), lung (n = 70) and endometrial (n = 87) cancer cells. A new pipeline is proposed integrating the DNA methylation intensity measurements on all the CpG islands by the transformation of Walsh-Hadamard Transform (WHT). The study reveals the three-step properties of the DNA methylation transform-domain data and the step values of association with the cell status. Further assessments have been carried out on the proposed machine learning pipeline to perform classification of the normal and cancer tissue cells. A number of machine learning classifiers are compared for whole sequence and WHT sequence classification based on public Whole-Genome Bisulfite Sequencing (WGBS) DNA methylation datasets. The WHT-based method can speed up the computation time by more than one order of magnitude compared with whole original sequence classification, while maintaining comparable classification accuracy by the selected machine learning classifiers. The proposed method has broad applications in expedited disease and normal human cell classifications by the epigenome and genome datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge