The interventional Bayesian Gaussian equivalent score for Bayesian causal inference with unknown soft interventions
Paper and Code
May 05, 2022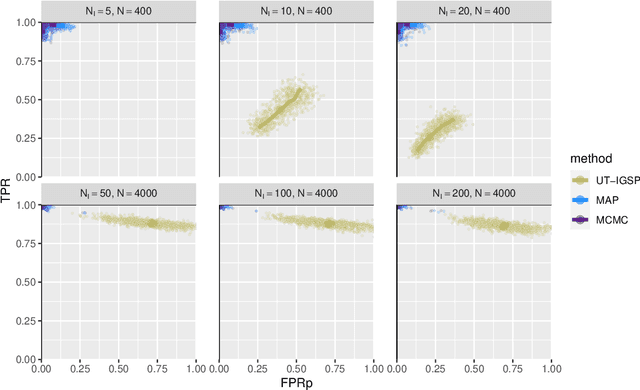

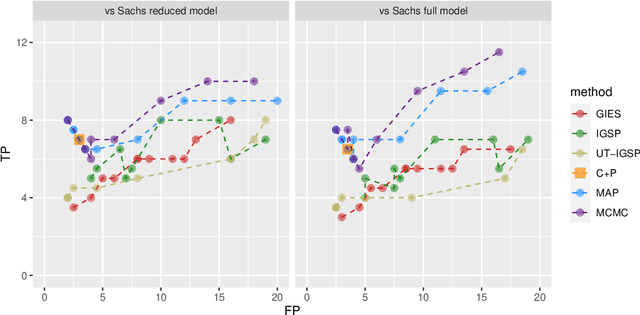

Describing the causal relations governing a system is a fundamental task in many scientific fields, ideally addressed by experimental studies. However, obtaining data under intervention scenarios may not always be feasible, while discovering causal relations from purely observational data is notoriously challenging. In certain settings, such as genomics, we may have data from heterogeneous study conditions, with soft (partial) interventions only pertaining to a subset of the study variables, whose effects and targets are possibly unknown. Combining data from experimental and observational studies offers the opportunity to leverage both domains and improve on the identifiability of causal structures. To this end, we define the interventional BGe score for a mixture of observational and interventional data, where the targets and effects of intervention may be unknown. To demonstrate the approach we compare its performance to other state-of-the-art algorithms, both in simulations and data analysis applications. Prerogative of our method is that it takes a Bayesian perspective leading to a full characterisation of the posterior distribution of the DAG structures. Given a sample of DAGs one can also automatically derive full posterior distributions of the intervention effects. Consequently the method effectively captures the uncertainty both in the structure and the parameter estimates. Codes to reproduce the simulations and analyses are publicly available at github.com/jackkuipers/iBGe
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge