NOISe: Nuclei-Aware Osteoclast Instance Segmentation for Mouse-to-Human Domain Transfer
Paper and Code
Apr 15, 2024
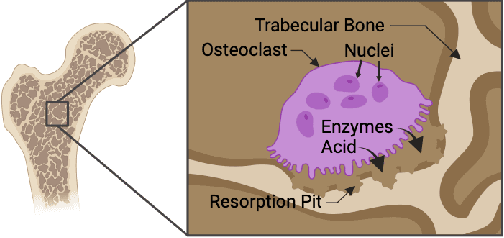
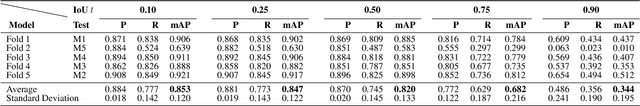
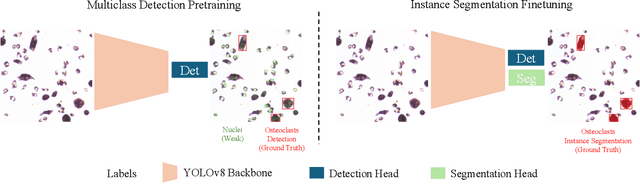
Osteoclast cell image analysis plays a key role in osteoporosis research, but it typically involves extensive manual image processing and hand annotations by a trained expert. In the last few years, a handful of machine learning approaches for osteoclast image analysis have been developed, but none have addressed the full instance segmentation task required to produce the same output as that of the human expert led process. Furthermore, none of the prior, fully automated algorithms have publicly available code, pretrained models, or annotated datasets, inhibiting reproduction and extension of their work. We present a new dataset with ~2*10^5 expert annotated mouse osteoclast masks, together with a deep learning instance segmentation method which works for both in vitro mouse osteoclast cells on plastic tissue culture plates and human osteoclast cells on bone chips. To our knowledge, this is the first work to automate the full osteoclast instance segmentation task. Our method achieves a performance of 0.82 mAP_0.5 (mean average precision at intersection-over-union threshold of 0.5) in cross validation for mouse osteoclasts. We present a novel nuclei-aware osteoclast instance segmentation training strategy (NOISe) based on the unique biology of osteoclasts, to improve the model's generalizability and boost the mAP_0.5 from 0.60 to 0.82 on human osteoclasts. We publish our annotated mouse osteoclast image dataset, instance segmentation models, and code at github.com/michaelwwan/noise to enable reproducibility and to provide a public tool to accelerate osteoporosis research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge