nD-PDPA: nDimensional Probability Density Profile Analysis
Paper and Code
Apr 05, 2023
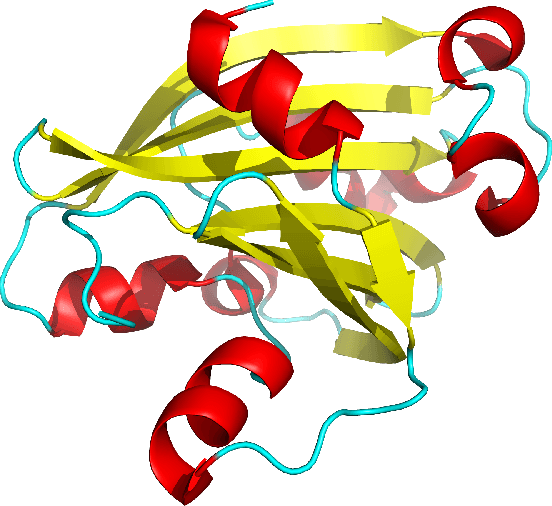
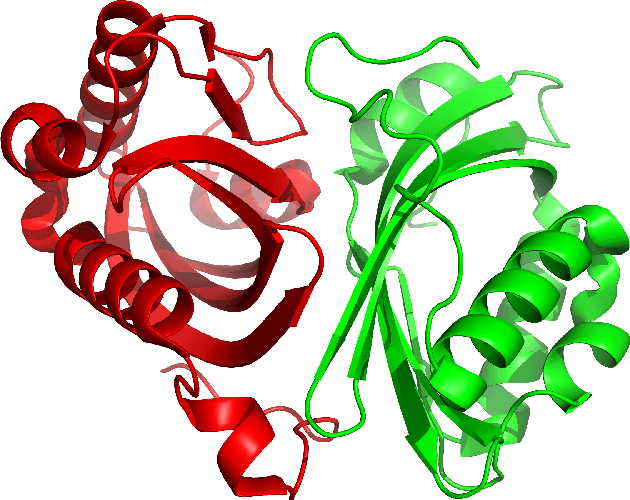
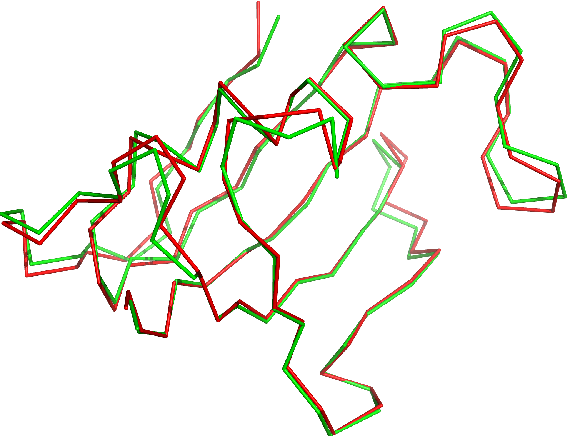
Despite the recent advances in various Structural Genomics Projects, a large gap remains between the number of sequenced and structurally characterized proteins. Some reasons for this discrepancy include technical difficulties, labor, and the cost related to determining a structure by experimental methods such as NMR spectroscopy. Several computational methods have been developed to expand the applicability of NMR spectroscopy by addressing temporal and economical problems more efficiently. While these methods demonstrate successful outcomes to solve more challenging and structurally novel proteins, the cost has not been reduced significantly. Probability Density Profile Analysis (PDPA) has been previously introduced by our lab to directly address the economics of structure determination of routine proteins and the identification of novel structures from a minimal set of unassigned NMR data. 2D-PDPA (in which 2D denotes incorporation of data from two alignment media) has been successful in identifying the structural homolog of an unknown protein within a library of ~1000 decoy structures. In order to further expand the selectivity and sensitivity of PDPA, the incorporation of additional data was necessary. However, the expansion of the original PDPA approach was limited by its computational requirements where the inclusion of additional data would render it computationally intractable. Here we present the most recent developments of PDPA method (nD-PDPA: n Dimensional Probability Density Profile Analysis) that eliminate 2D-PDPA's computational limitations, and allows inclusion of RDC data from multiple vector types in multiple alignment media.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge