Molecular Hypergraph Grammar with its Application to Molecular Optimization
Paper and Code
Sep 08, 2018

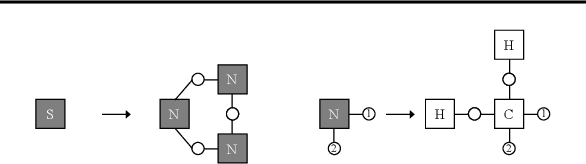

This paper is concerned with a molecular optimization framework using variational autoencoders (VAEs). In this paradigm, VAE allows us to convert a molecular graph into/from its latent continuous vector, and therefore, the molecular optimization problem can be solved by continuous optimization techniques. One of the longstanding issues in this area is that it is difficult to always generate valid molecules. The very recent work called the junction tree variational autoencoder (JT-VAE) successfully solved this issue by generating a molecule fragment-by-fragment. While it achieves the state-of-the-art performance, it requires several neural networks to be trained, which predict which atoms are used to connect fragments and stereochemistry of each bond. In this paper, we present a molecular hypergraph grammar variational autoencoder (MHG-VAE), which uses a single VAE to address the issue. Our idea is to develop a novel graph grammar for molecular graphs called molecular hypergraph grammar (MHG), which can specify the connections between fragments and the stereochemistry on behalf of neural networks. This capability allows us to address the issue using only a single VAE. We empirically demonstrate the effectiveness of MHG-VAE over existing methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge