Graph Neural Network for Cell Tracking in Microscopy Videos
Paper and Code
Feb 09, 2022
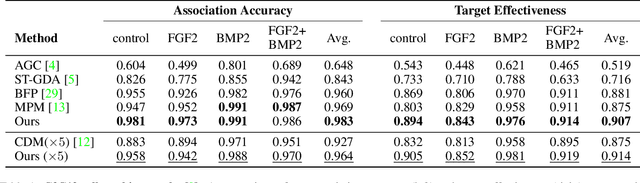

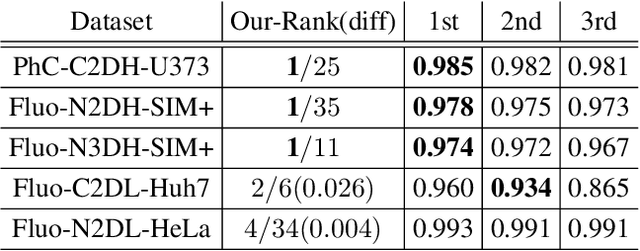
We present a novel graph neural network (GNN) approach for cell tracking in high-throughput microscopy videos. By modeling the entire time-lapse sequence as a direct graph where cell instances are represented by its nodes and their associations by its edges, we extract the entire set of cell trajectories by looking for the maximal paths in the graph. This is accomplished by several key contributions incorporated into an end-to-end deep learning framework. We exploit a deep metric learning algorithm to extract cell feature vectors that distinguish between instances of different biological cells and assemble same cell instances. We introduce a new GNN block type which enables a mutual update of node and edge feature vectors, thus facilitating the underlying message passing process. The message passing concept, whose extent is determined by the number of GNN blocks, is of fundamental importance as it enables the `flow' of information between nodes and edges much behind their neighbors in consecutive frames. Finally, we solve an edge classification problem and use the identified active edges to construct the cells' tracks and lineage trees. We demonstrate the strengths of the proposed cell tracking approach by applying it to 2D and 3D datasets of different cell types, imaging setups, and experimental conditions. We show that our framework outperforms most of the current state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge