Fast marginal likelihood estimation of penalties for group-adaptive elastic net
Paper and Code
Jan 11, 2021
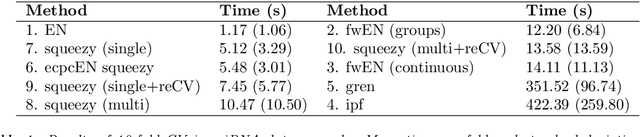
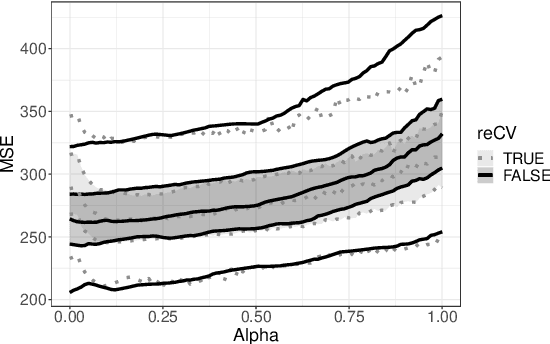
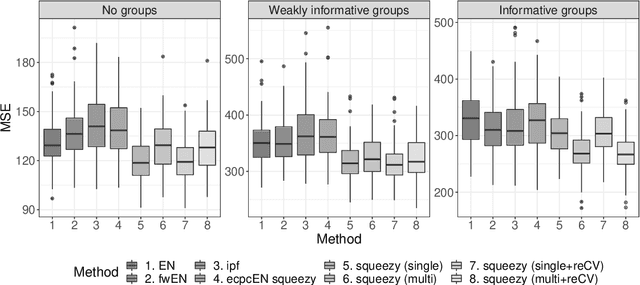
Nowadays, clinical research routinely uses omics data, such as gene expression, for predicting clinical outcomes or selecting markers. Additionally, so-called co-data are often available, providing complementary information on the covariates, like p-values from previously published studies or groups of genes corresponding to pathways. Elastic net penalisation is widely used for prediction and covariate selection. Group-adaptive elastic net penalisation learns from co-data to improve the prediction and covariate selection, by penalising important groups of covariates less than other groups. Existing methods are, however, computationally expensive. Here we present a fast method for marginal likelihood estimation of group-adaptive elastic net penalties for generalised linear models. We first derive a low-dimensional representation of the Taylor approximation of the marginal likelihood and its first derivative for group-adaptive ridge penalties, to efficiently estimate these penalties. Then we show by using asymptotic normality of the linear predictors that the marginal likelihood for elastic net models may be approximated well by the marginal likelihood for ridge models. The ridge group penalties are then transformed to elastic net group penalties by using the variance function. The method allows for overlapping groups and unpenalised variables. We demonstrate the method in a model-based simulation study and an application to cancer genomics. The method substantially decreases computation time and outperforms or matches other methods by learning from co-data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge