Dependency detection with similarity constraints
Paper and Code
Jan 31, 2011
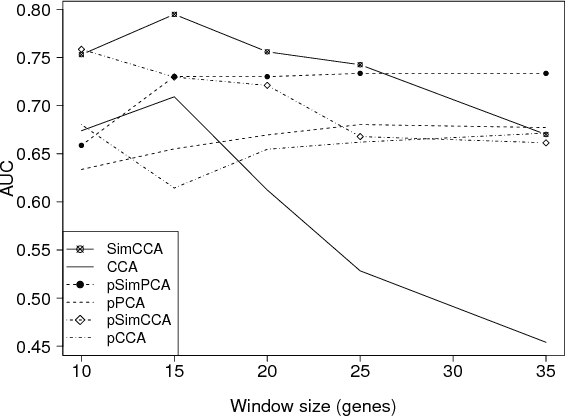
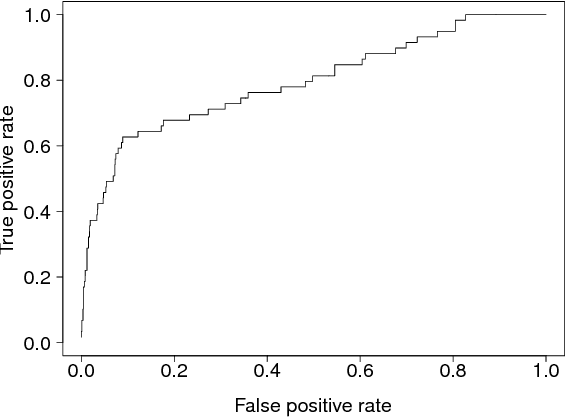
Unsupervised two-view learning, or detection of dependencies between two paired data sets, is typically done by some variant of canonical correlation analysis (CCA). CCA searches for a linear projection for each view, such that the correlations between the projections are maximized. The solution is invariant to any linear transformation of either or both of the views; for tasks with small sample size such flexibility implies overfitting, which is even worse for more flexible nonparametric or kernel-based dependency discovery methods. We develop variants which reduce the degrees of freedom by assuming constraints on similarity of the projections in the two views. A particular example is provided by a cancer gene discovery application where chromosomal distance affects the dependencies between gene copy number and activity levels. Similarity constraints are shown to improve detection performance of known cancer genes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge