Context-Aware Self-Supervised Learning of Whole Slide Images
Paper and Code
Jun 07, 2023

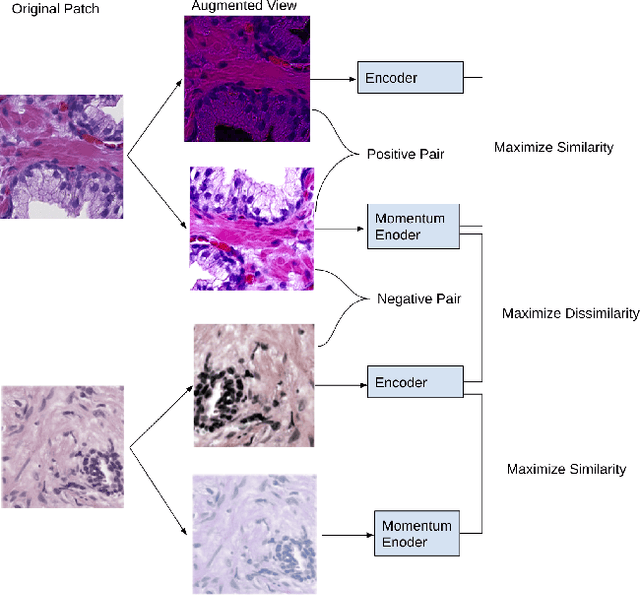

Presenting whole slide images (WSIs) as graph will enable a more efficient and accurate learning framework for cancer diagnosis. Due to the fact that a single WSI consists of billions of pixels and there is a lack of vast annotated datasets required for computational pathology, the problem of learning from WSIs using typical deep learning approaches such as convolutional neural network (CNN) is challenging. Additionally, WSIs down-sampling may lead to the loss of data that is essential for cancer detection. A novel two-stage learning technique is presented in this work. Since context, such as topological features in the tumor surroundings, may hold important information for cancer grading and diagnosis, a graph representation capturing all dependencies among regions in the WSI is very intuitive. Graph convolutional network (GCN) is deployed to include context from the tumor and adjacent tissues, and self-supervised learning is used to enhance training through unlabeled data. More specifically, the entire slide is presented as a graph, where the nodes correspond to the patches from the WSI. The proposed framework is then tested using WSIs from prostate and kidney cancers. To assess the performance improvement through self-supervised mechanism, the proposed context-aware model is tested with and without use of pre-trained self-supervised layer. The overall model is also compared with multi-instance learning (MIL) based and other existing approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge