A generative recommender system with GMM prior for cancer drug generation and sensitivity prediction
Paper and Code
Jun 07, 2022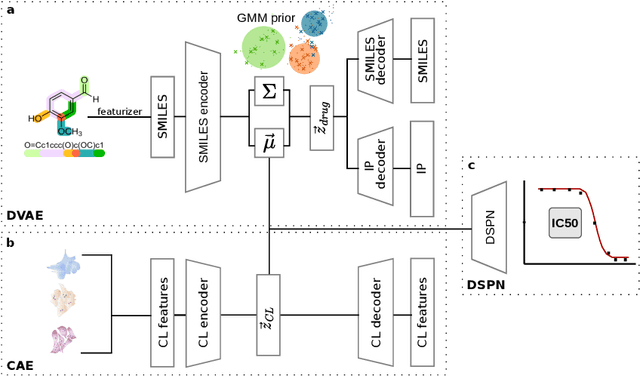
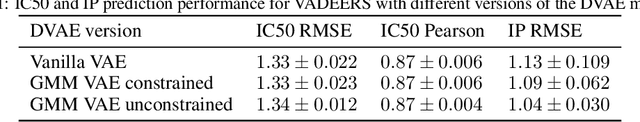

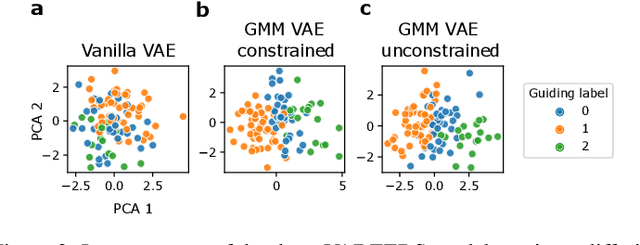
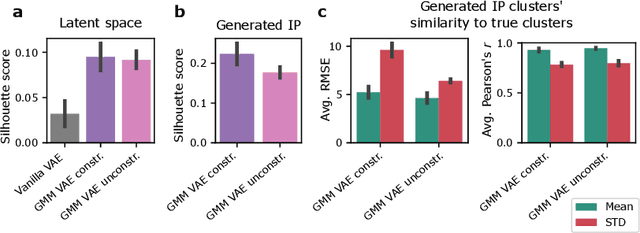
Recent emergence of high-throughput drug screening assays sparkled an intensive development of machine learning methods, including models for prediction of sensitivity of cancer cell lines to anti-cancer drugs, as well as methods for generation of potential drug candidates. However, a concept of generation of compounds with specific properties and simultaneous modeling of their efficacy against cancer cell lines has not been comprehensively explored. To address this need, we present VADEERS, a Variational Autoencoder-based Drug Efficacy Estimation Recommender System. The generation of compounds is performed by a novel variational autoencoder with a semi-supervised Gaussian Mixture Model (GMM) prior. The prior defines a clustering in the latent space, where the clusters are associated with specific drug properties. In addition, VADEERS is equipped with a cell line autoencoder and a sensitivity prediction network. The model combines data for SMILES string representations of anti-cancer drugs, their inhibition profiles against a panel of protein kinases, cell lines biological features and measurements of the sensitivity of the cell lines to the drugs. The evaluated variants of VADEERS achieve a high r=0.87 Pearson correlation between true and predicted drug sensitivity estimates. We train the GMM prior in such a way that the clusters in the latent space correspond to a pre-computed clustering of the drugs by their inhibitory profiles. We show that the learned latent representations and new generated data points accurately reflect the given clustering. In summary, VADEERS offers a comprehensive model of drugs and cell lines properties and relationships between them, as well as a guided generation of novel compounds.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge