A bi-atrial statistical shape model for large-scale in silico studies of human atria: model development and application to ECG simulations
Paper and Code
Feb 22, 2021
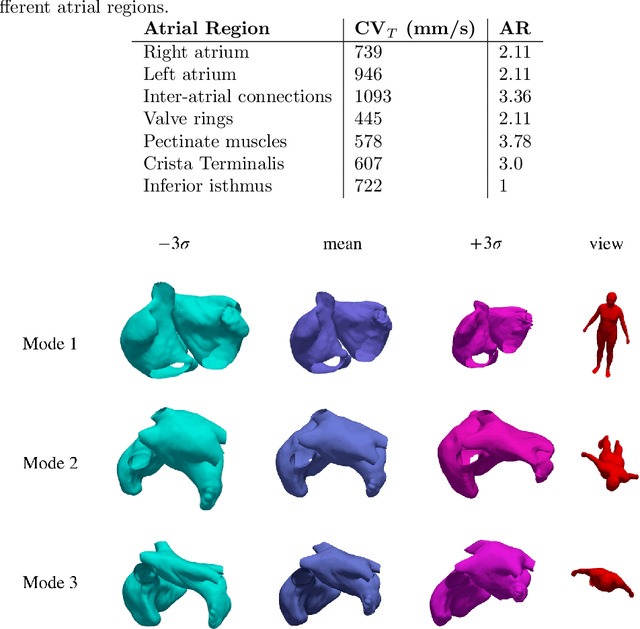
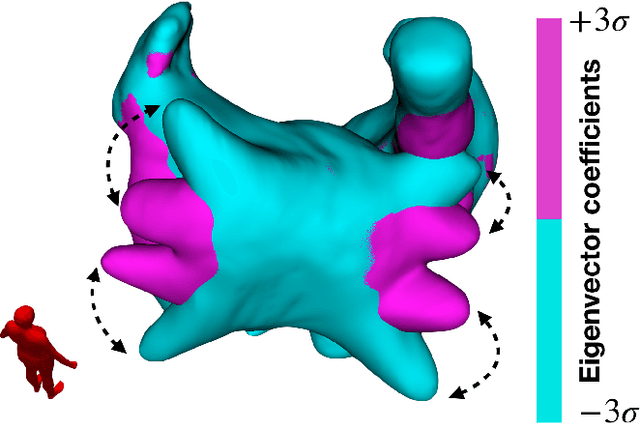

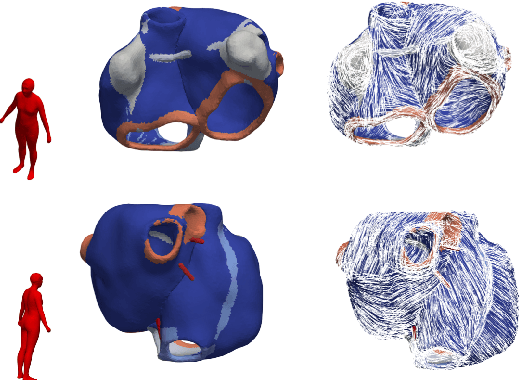
Large-scale electrophysiological simulations to obtain electrocardiograms (ECG) carry the potential to produce extensive datasets for training of machine learning classifiers to, e.g., discriminate between different cardiac pathologies. The adoption of simulations for these purposes is limited due to a lack of ready-to-use models covering atrial anatomical variability. We built a bi-atrial statistical shape model (SSM) of the endocardial wall based on 47 segmented human CT and MRI datasets using Gaussian process morphable models. Generalization, specificity, and compactness metrics were evaluated. The SSM was applied to simulate atrial ECGs in 100 random volumetric instances. The first eigenmode of our SSM reflects a change of the total volume of both atria, the second the asymmetry between left vs. right atrial volume, the third a change in the prominence of the atrial appendages. The SSM is capable of generalizing well to unseen geometries and 95% of the total shape variance is covered by its first 23 eigenvectors. The P waves in the 12-lead ECG of 100 random instances showed a duration of 104ms in accordance with large cohort studies. The novel bi-atrial SSM itself as well as 100 exemplary instances with rule-based augmentation of atrial wall thickness, fiber orientation, inter-atrial bridges and tags for anatomical structures have been made publicly available. The novel, openly available bi-atrial SSM can in future be employed to generate large sets of realistic atrial geometries as a basis for in silico big data approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge