Sophie Peacock
Coupling Deep Imputation with Multitask Learning for Downstream Tasks on Genomics Data
Apr 28, 2022
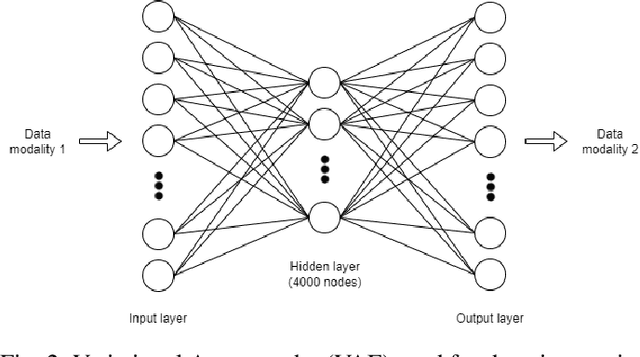

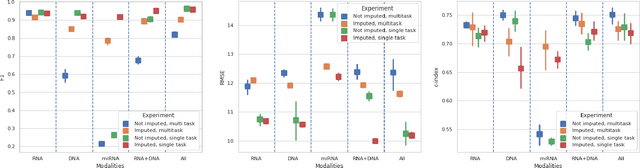
Abstract:Genomics data such as RNA gene expression, methylation and micro RNA expression are valuable sources of information for various clinical predictive tasks. For example, predicting survival outcomes, cancer histology type and other patients' related information is possible using not only clinical data but molecular data as well. Moreover, using these data sources together, for example in multitask learning, can boost the performance. However, in practice, there are many missing data points which leads to significantly lower patient numbers when analysing full cases, which in our setting refers to all modalities being present. In this paper we investigate how imputing data with missing values using deep learning coupled with multitask learning can help to reach state-of-the-art performance results using combined genomics modalities, RNA, micro RNA and methylation. We propose a generalised deep imputation method to impute values where a patient has all modalities present except one. Interestingly enough, deep imputation alone outperforms multitask learning alone for the classification and regression tasks across most combinations of modalities. In contrast, when using all modalities for survival prediction we observe that multitask learning alone outperforms deep imputation alone with statistical significance (adjusted p-value 0.03). Thus, both approaches are complementary when optimising performance for downstream predictive tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge